Table of Contents
- Предисловие
- Введение
- Выборка отождествлений галактик из RFGC
- Этапы «очистки»
- 1. Выберем всё, что внутри эллипса, заданного RFGC и круга в [Math Processing Error]$10'$ от положения центра в RFGC
- 2. Оставим только те, которые есть в
ForcedGalaxyShapeв заданной полосе - 3. Оставим только те, для которых вписан Серсик в заданной полосе
- 4. Отсеиваем программные ошибки вписания профиля
- 5. Выбор согласованных по позиционному углу источников
- Выбор единственного наилучшего отождествления
- Этапы «очистки»
- Подбор критериев отбора (g)
- Подбор критериев отбора (r)
- Подбор критериев отбора (i)
- Подбор критериев отбора (z)
- Подбор критериев отбора (y)
- Отбор по выполнению критерия в любой полосе
Предисловие¶
Glossary¶
Term | Explanation |
|---|---|
Term | Explanation |
| Detection | Catalog data for a single source from a single exposure. |
| Object | Catalog data for one or more associated detections. |
| OTA | Orthogonal Transfer Array. An OTA consists of an 8 x 8 array of 600 x 600 pixel CCD cells. The PS1 camera consists of 60 OTAs. |
| PSPS | Published Science Product System. This is the PS1 object database. |
| Sky cell | One tile in the PS1 tessellation of the sky. |
| Stack image | Image constructed by combining all useful warp images that overlap a single sky cell. Sometimes simply referred to as a "stack". |
| Stack detection | Catalog data for a single source from a stack image. |
| Stack object | Catalog data for one or more associated stack detections. |
| Tessellation | The scheme used to cover the spherical sky with a regular pattern of rectangular images in a manner that leaves no gaps in coverage. |
| Warp image | Image constructed by resampling a single exposure onto a canonical sky cell projection. Sometimes simply referred to as a "warp". |
| Ubercal | The process that uses comparisons in flux measurements from overlapping exposures to improve the global calibration. |
Рабочие полосы¶
| band | lambda | spec.res |
|---|---|---|
| g | 481 nm | (R = 3.5) |
| r | 617 nm | (R = 4.4) |
| i | 752 nm | (R = 5.8) |
| z | 866 nm | (R = 8.3) |
| y | 962 nm | (R = 11.6) |
Советы про использование звездных величин¶
- For point sources use PSF magnitudes.
- Mean PSF magnitudes have the lowest noise (because the PSF model is most accurate in single-epoch images). They are good for brighter objects, but for objects near the single-epoch detection limit they will be biased (due to the absence of sub-threshold detections), and objects too faint to detect in a single epoch are missing.
- Stack PSF magnitudes are noisier because the PSF model is less accurate. But the stack detections are more than a magnitude deeper and so have many more faint objects than mean detections.
- Forced mean PSF magnitudes use PSF-fitting photometry on the single-epoch images at positions of stack detections. They are a reasonable compromise: they have slightly lower noise than the stack PSF magnitudes, and they are deep and unbiased (because they use data from all warps). Their noise is higher than the mean PSF magnitudes, however.
- For extended objects use Kron magnitudes.
- Stack Kron magnitudes are usually the first choice as a general-purpose, deep magnitude.
- de Vaucouleurs and exponential model fits could be better in some cases, and the mean measurements can be useful for objects that are barely resolved (where the PSF is important).
- Extended object photometry using the PS1 catalog will require research and analysis by the user to determine the best approach.
- For point sources use PSF magnitudes.
Представляющие интерес параметры и таблицы¶
MeanObjectView=MeanObjectJOINObjectThin-- фотометрия (PSF, Kron, апертурная) и координаты. Отсюда брали:raMean,decMean{filt}MeanKronMag
StackModelFitSer-- вписанный Серсик в изображение в разных полосах. Профиль Серсика у них такой (если убрать перекрестные члены): $$ \begin{aligned} \rho &= \sqrt{x^2/R_{xx}^2 + y^2/R_{yy}^2} \\ f &= I_0 e^{-\rho ^{1/n}} \end{aligned} $$ Отсюда брали:{filt}SerRadius-- какой-то параметр профиля, скорее всего $R_{xx}${filt}SerAb--- отношение $b/a$ (а не $a/b$){filt}SerNu-- индекс $n${filt}SerPhi-- позиционный угол. Отсчитывается от направления на восток (почему-то)
ForcedGalaxyShape-- сюдя по всему, сначала изображение разбивается на 24 сектора, потом в каждом строится гистограмма, ищется медиана, и уже по этим точкам вписывается эллипс, но это неточно. Отсюда брали:{filt}GalMajor-- большая полуось вписанного эллипса{filt}GalMinor--- малая полуось вписанного эллипса{filt}GalPhi-- позиционный угол. Отсчитывается от направления на восток (почему-то)
подробнее -- тут
Статьи¶
- The Pan-STARRS1 Surveys, Chambers, K.C., et al.
- Pan-STARRS Data Processing System, Magnier, E. A., et al.
- Pan-STARRS Pixel Processing: Detrending, Warping, Stacking, Waters, C. Z., et al.
- Pan-STARRS Pixel Analysis: Source Detection and Characterization, Magnier, E. A., et al.
- Pan-STARRS Photometric and Astrometric Calibration, Magnier, E. A., et al.
- The Pan-STARRS1 Database and Data Products, Flewelling, H. A., et al.
Про тонкости того, что означают те или иные параметры стоит идти в [4], но написано очень непонятно
Введение¶
Основная цель --- научиться выбирать плоские галактики в PanSTARRS. Судя по всему, они же будут видимыми с ребра, но для этого нужно анализировать вертикальный профиль, а такую задачу пока не пытались решать
Задачи:
- Сопоставить какие-то объекты в PanSTARRS объектам из RFGC
- Придумать критерий отбора плоских галактик, анализируя эти сопоставления
- Протестировать эти критерии и отобрать кандидаты
Выборка отождествлений галактик из RFGC¶
Запрос чтобы достать все объекты в пределах заданного радиуса ($10''$) во всех полосах
WITH
nearobj AS (
SELECT
nb.objID, r.RFGC
FROM
MyDB.RFGCfull AS r
CROSS APPLY fGetNearbyObjEq(r.RAJ2000, r.DEJ2000, 0.166) AS nb
-- WHERE r.RFGC < 10
)
, panpos AS (
SELECT
o.objID, o.objName, o.raMean, o.decMean
FROM
MeanObjectView AS o
INNER JOIN nearobj ON (o.objID=nearobj.objID)
)
, ser AS (
SELECT s.*
FROM
StackModelFitSer AS s
INNER JOIN nearobj ON (s.objID=nearobj.objID)
)
, seruniq AS (
SELECT *
FROM
( SELECT *
,ROW_NUMBER () OVER (
PARTITION BY objID
ORDER BY bestDetection DESC, primaryDetection DESC
) AS rown -- enumerate inside common id group
FROM ser
) AS t
WHERE t.rown = 1 -- only first
)
, shape AS (
SELECT s.*
FROM ForcedGalaxyShape AS s
INNER JOIN nearobj ON (s.objID=nearobj.objID)
)
, shapeuniq AS (
SELECT *
FROM (
SELECT
*
, ROW_NUMBER () OVER (
PARTITION BY objID
ORDER BY
CASE WHEN gGalFlags = 0 THEN 1
WHEN gGalFlags = 4 THEN 2
ELSE 3 END ASC,
CASE WHEN rGalFlags = 0 THEN 1
WHEN rGalFlags = 4 THEN 2
ELSE 3 END ASC,
CASE WHEN iGalFlags = 0 THEN 1
WHEN iGalFlags = 4 THEN 2
ELSE 3 END ASC,
CASE WHEN zGalFlags = 0 THEN 1
WHEN zGalFlags = 4 THEN 2
ELSE 3 END ASC,
CASE WHEN yGalFlags = 0 THEN 1
WHEN yGalFlags = 4 THEN 2
ELSE 3 END ASC
) AS rown -- enumerate inside common id group
FROM shape
) AS t
WHERE t.rown = 1 -- only first
)
, pet AS (
SELECT p.*
FROM
StackPetrosian AS p
INNER JOIN nearobj ON (p.objID=nearobj.objID)
)
, petuniq AS (
SELECT *
FROM
( SELECT *
,ROW_NUMBER () OVER (
PARTITION BY objID
ORDER BY bestDetection DESC, primaryDetection DESC
) AS rown -- enumerate inside common id group
FROM pet
) AS t
WHERE t.rown = 1 -- only first
)
, kron AS (
SELECT
a.objID,
a.bestDetection AS RadiusbestDetection, a.primaryDetection AS RadiusprimaryDetection,
o.gMeanKronMag AS gkronMag, a.gKronRad AS gkronRadius,
o.rMeanKronMag AS rkronMag, a.rKronRad AS rkronRadius,
o.iMeanKronMag AS ikronMag, a.iKronRad AS ikronRadius,
o.zMeanKronMag AS zkronMag, a.zKronRad AS zkronRadius,
o.yMeanKronMag AS ykronMag, a.yKronRad AS ykronRadius
FROM
StackObjectAttributes AS a
INNER JOIN nearobj ON (a.objID=nearobj.objID)
LEFT JOIN MeanObjectView AS o on (o.objID=a.objID)
)
, kronuniq AS (
SELECT *
FROM
( SELECT *
,ROW_NUMBER () OVER (
PARTITION BY objID
ORDER BY RadiusbestDetection DESC, RadiusprimaryDetection DESC
) AS rown -- enumerate inside common id group
FROM kron
) AS t
WHERE t.rown = 1 -- only first
)
SELECT
rfgc.*,
panpos.objID,
panpos.objName,
panpos.raMean,
panpos.decMean
,
seruniq.bestDetection AS serbestDetection,
seruniq.primaryDetection AS serprimaryDetection
,
seruniq.gSerRadius,seruniq.gSerAb,seruniq.gSerPhi,seruniq.gSerRa,seruniq.gSerDec,seruniq.gSerMag,
seruniq.rSerRadius,seruniq.rSerAb,seruniq.rSerPhi,seruniq.rSerRa,seruniq.rSerDec,seruniq.rSerMag,
seruniq.iSerRadius,seruniq.iSerAb,seruniq.iSerPhi,seruniq.iSerRa,seruniq.iSerDec,seruniq.iSerMag,
seruniq.zSerRadius,seruniq.zSerAb,seruniq.zSerPhi,seruniq.zSerRa,seruniq.zSerDec,seruniq.zSerMag,
seruniq.ySerRadius,seruniq.ySerAb,seruniq.ySerPhi,seruniq.ySerRa,seruniq.ySerDec,seruniq.ySerMag
,
shapeuniq.gGalMinor,shapeuniq.gGalMajor,shapeuniq.gGalPhi,shapeuniq.gGalIndex,shapeuniq.gGalMag,shapeuniq.gGalFlags,
shapeuniq.rGalMinor,shapeuniq.rGalMajor,shapeuniq.rGalPhi,shapeuniq.rGalIndex,shapeuniq.rGalMag,shapeuniq.rGalFlags,
shapeuniq.iGalMinor,shapeuniq.iGalMajor,shapeuniq.iGalPhi,shapeuniq.iGalIndex,shapeuniq.iGalMag,shapeuniq.iGalFlags,
shapeuniq.zGalMinor,shapeuniq.zGalMajor,shapeuniq.zGalPhi,shapeuniq.zGalIndex,shapeuniq.zGalMag,shapeuniq.zGalFlags,
shapeuniq.yGalMinor,shapeuniq.yGalMajor,shapeuniq.yGalPhi,shapeuniq.yGalIndex,shapeuniq.yGalMag,shapeuniq.yGalFlags
,
petuniq.bestDetection AS petbestDetection,
petuniq.primaryDetection AS petprimaryDetection
,
kronuniq.RadiusbestDetection AS kronRadiusbestDetection,
kronuniq.RadiusprimaryDetection AS kronRadiusprimaryDetection
,
petuniq.gpetMag,petuniq.gpetRadius,petuniq.rpetMag,petuniq.rpetRadius,petuniq.ipetMag,petuniq.ipetRadius,petuniq.zpetMag,petuniq.zpetRadius,petuniq.ypetMag,petuniq.ypetRadius,
kronuniq.gkronMag,kronuniq.gkronRadius,kronuniq.rkronMag,kronuniq.rkronRadius,kronuniq.ikronMag,kronuniq.ikronRadius,kronuniq.zkronMag,kronuniq.zkronRadius,kronuniq.ykronMag,kronuniq.ykronRadius
INTO rfgc_nearby_multiband2
FROM MyDB.RFGCfull as rfgc
LEFT JOIN nearobj ON nearobj.RFGC = rfgc.RFGC
LEFT JOIN panpos ON panpos.objID = nearobj.objID
LEFT JOIN seruniq ON seruniq.objID = panpos.objID
LEFT JOIN shapeuniq ON shapeuniq.objID = seruniq.objID
LEFT JOIN kronuniq ON kronuniq.objID = shapeuniq.objID
LEFT JOIN petuniq ON petuniq.objID = kronuniq.objID
Здесь *uniq -- таблицы в которых выбрано «лучшее» детектирование для заданного objID
import numpy as np
import yaml
import numpy.ma as ma
from argparse import Namespace
import hashlib
import re
from IPython.display import display, HTML
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
%load_ext autoreload
%autoreload 2
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import SymLogNorm
import matplotlib as mpl
# строить картинки высокого разрешения
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.image as mpimg
from astropy.visualization import LogStretch, PercentileInterval
import plotly.graph_objects as go
import plotly.io as pio
import plotly.tools as plyt
import plotly.express as px
import plotly.figure_factory as ff
from plotly.subplots import make_subplots
png_renderer = pio.renderers["png"]
png_renderer.width = 800
png_renderer.height = 600
png_renderer.scale = 3
pio.renderers.default = "png"
pio.templates.default = "plotly_white"
#pio.renderers.default = 'png+jupyterlab'
from astropy.modeling import fitting, models
from astropy.coordinates import SkyCoord, Angle
import astropy.units as u
from joblib import Memory
cachedir = "./cached/"
memory = Memory(cachedir, verbose=0)
import pandas as pd
def mw_omit(l):
b0 = 20 # deg
rb = 15 # deg
sigmab = 50 # deg
lw = Angle(l, unit=u.deg).wrap_at('180d').value
return b0 + rb*np.exp(-lw**2/(2*sigmab**2))
N = int(1e5)
points = np.random.uniform(-1, 1, size=(N, 3))
points = points[np.linalg.norm(points, axis=1) < 1]
coords = SkyCoord(points, representation_type='cartesian')
galaxies = pd.DataFrame({
'ra' : coords.cirs.ra,
'dec' : coords.cirs.dec,
'l' : coords.galactic.l,
'b' : coords.galactic.b
})
galaxies_sel = galaxies[
(np.abs(galaxies.b) > mw_omit(galaxies.l)) &
(galaxies.dec > -30)
]
gal_disc_ratio = len(galaxies_sel)/len(galaxies)
import seaborn as sb
# настройка стиля
sb.set(rc={'figure.figsize':(4,3)})
sb.set_style('whitegrid', {'grid.linestyle': ':'})
sb.set_palette("bright")
# ещё более тонкая настройка отображения картинок
dpi = 150
plt.rc('text', usetex=False)
plt.rc("savefig", dpi=dpi)
plt.rc("figure", dpi=dpi)
plt.rc('xtick', direction='in')
plt.rc('ytick', direction='in')
SMALL_SIZE = 8
MEDIUM_SIZE = 10
BIGGER_SIZE = 12
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
#plt.rc('xtick.major', pad=5)
#plt.rc('xtick.minor', pad=5)
#plt.rc('ytick.major', pad=5)
#plt.rc('ytick.minor', pad=5)
plt.rc('lines', dotted_pattern = [0.5, 1.1])
plt.rc('axes', axisbelow=False)
from astropy.table import Table
from astropy.coordinates import Angle
import astropy.units as u
from astropy.io import fits
from astropy.wcs import WCS
from pathlib import Path
datapath = Path('data')
querypath = Path('queries')
name = f"rfgc_nearby_multiband2"
bands = 'grizy'
df = pd.read_hdf(datapath / (name + ".h5"))
#with pd.option_context("display.max_rows", 1000):
# display(df[1:3].T)
for f in bands:
df[f"{f}SerA"] = df[f"{f}SerRadius"]
df[f"{f}SerB"] = df[f"{f}SerA"] * df[f"{f}SerAb"]
df[f"{f}SerBa"] = 1/df[f"{f}SerAb"]
df[f"{f}GalA"] = df[f"{f}GalMajor"]
df[f"{f}GalB"] = df[f"{f}GalMinor"]
df[f"{f}GalAb"] = df[f"{f}GalMinor"] / df[f"{f}GalMajor"]
df[f"{f}GalBa"] = 1/df[f"{f}GalAb"]
df["AbO"] = df['bO'] / df['aO']
При таком выборе получается много отождествлений на один объект RFGC
ids_per_rfgc = df.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
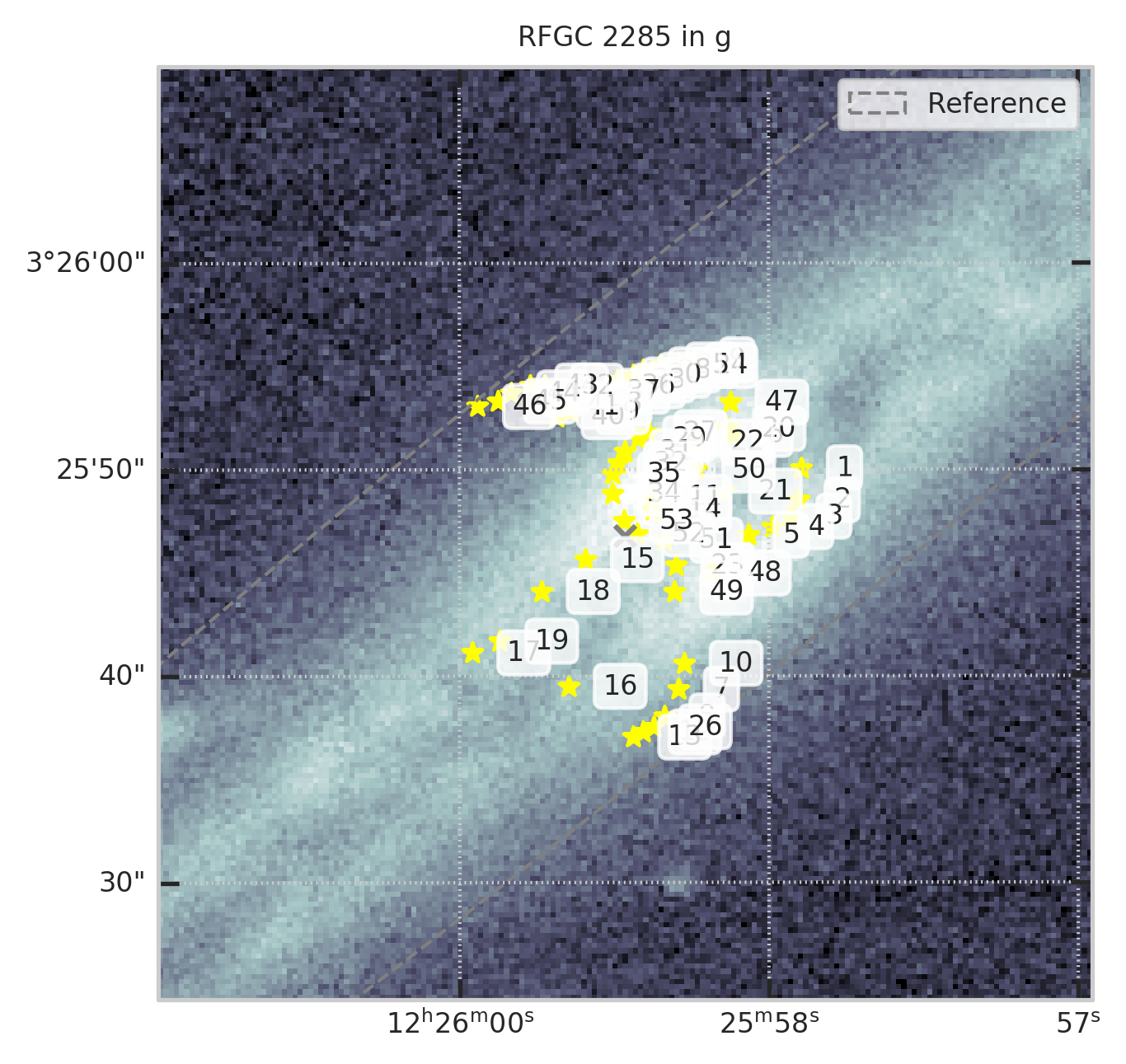
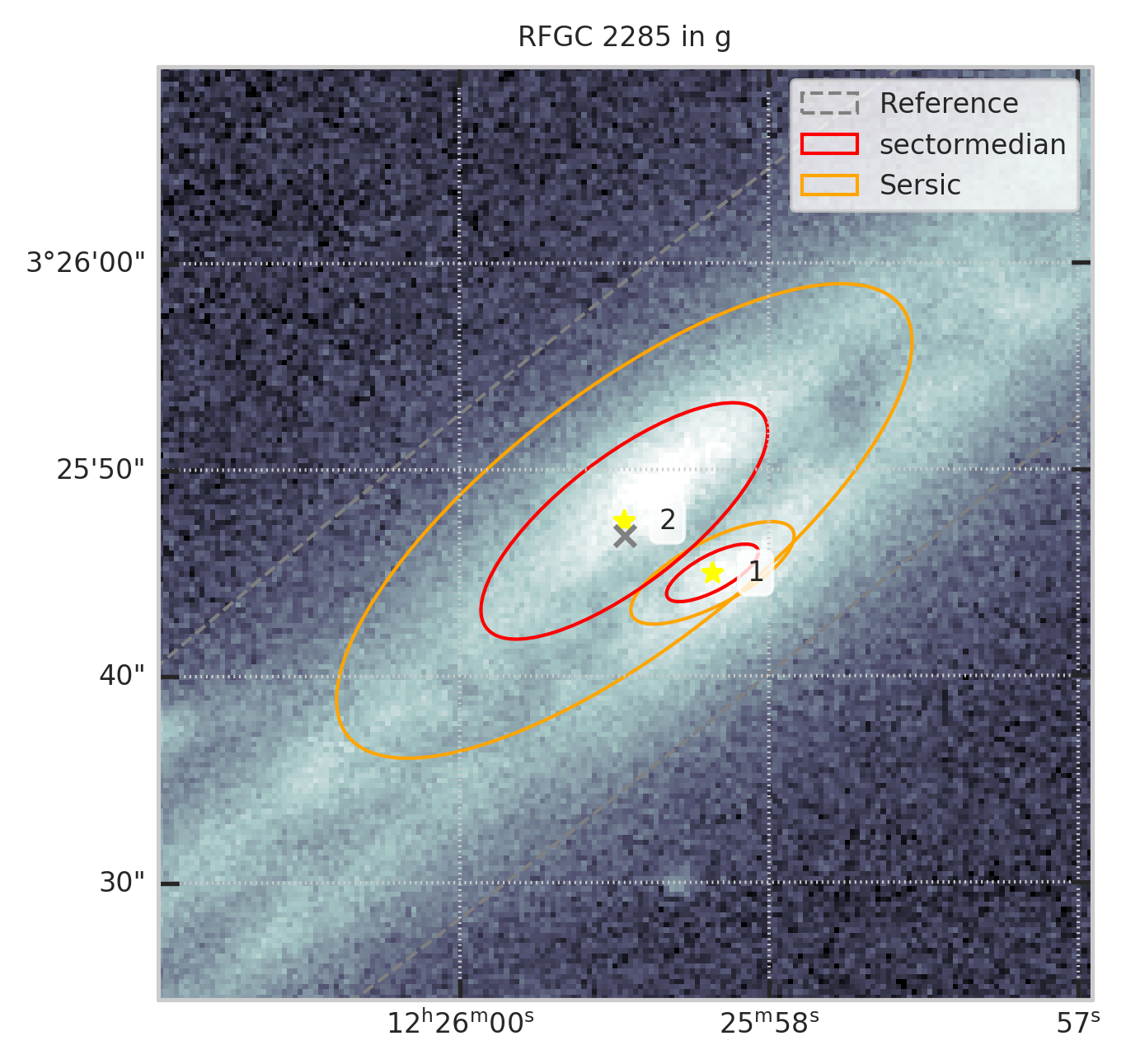
Например, для RFGC 2285 в полосе g
from code.crosstools import show_galaxy_rfgc
fig = plt.figure(figsize=(5,5))
subset = df[df.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=False, cleanp=False, kron=False, median=False, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
Этапы «очистк軶
1. Выберем всё, что внутри эллипса, заданного RFGC и круга в $10'$ от положения центра в RFGC¶
from code.crosstools import inellipse
ra, dec = df['raMean'], df['decMean']
ra_ref, dec_ref = df['RAJ2000'], df['DEJ2000']
sigma_a = np.minimum(df['aO'], 10.)
sigma_b = np.minimum(df['bO'], 10.)
mask = inellipse((ra, dec), (ra_ref, dec_ref), -df['PA'], sigma_a, sigma_b)
matches_ref = df.loc[mask]
fig = plt.figure(figsize=(5,5))
t = matches_ref
subset = t[t.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=False, cleanp=False, kron=False, median=False, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
print(f"Осталось ~{len(t)/len(df)*100:.0f}% объектов")
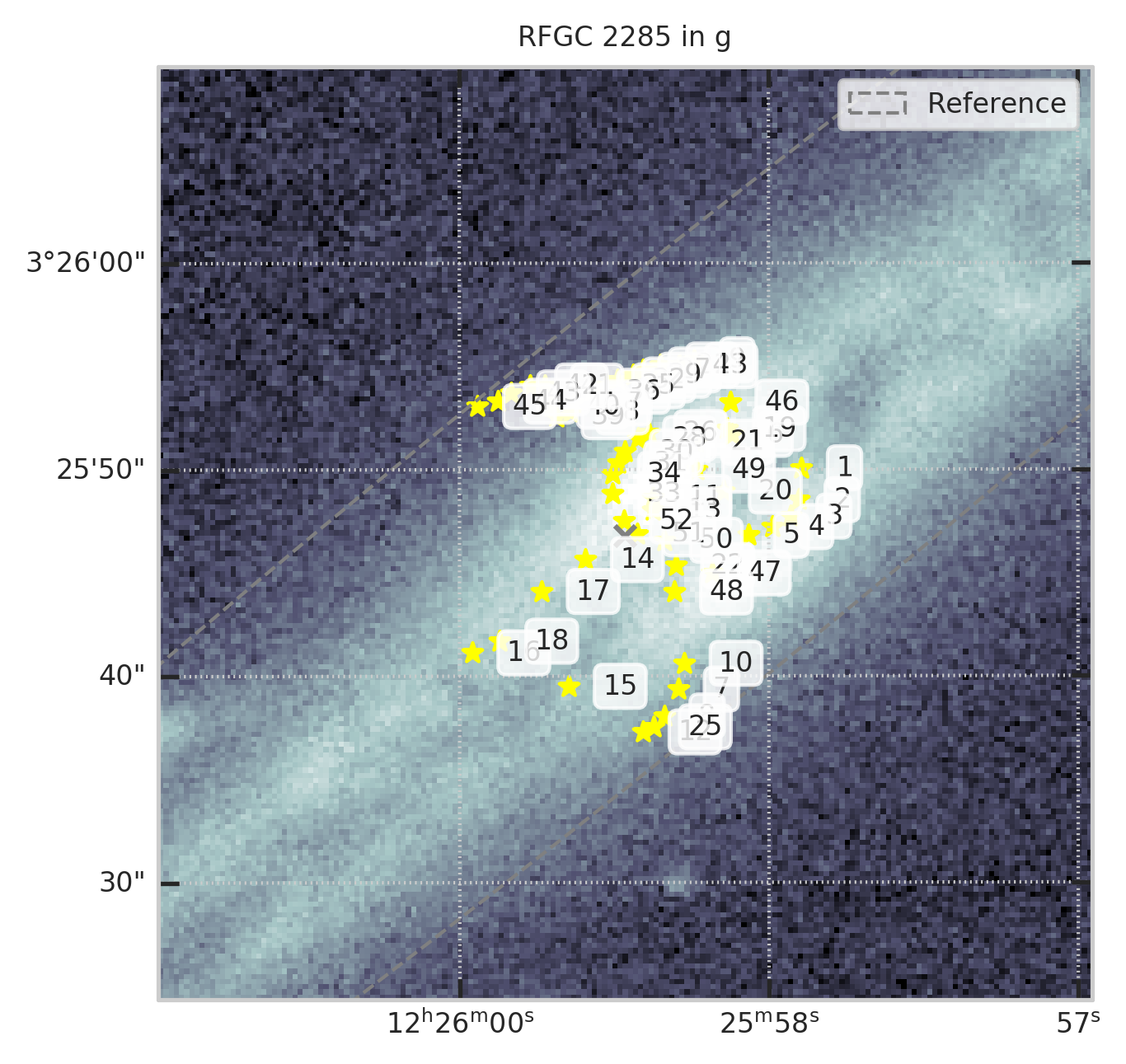
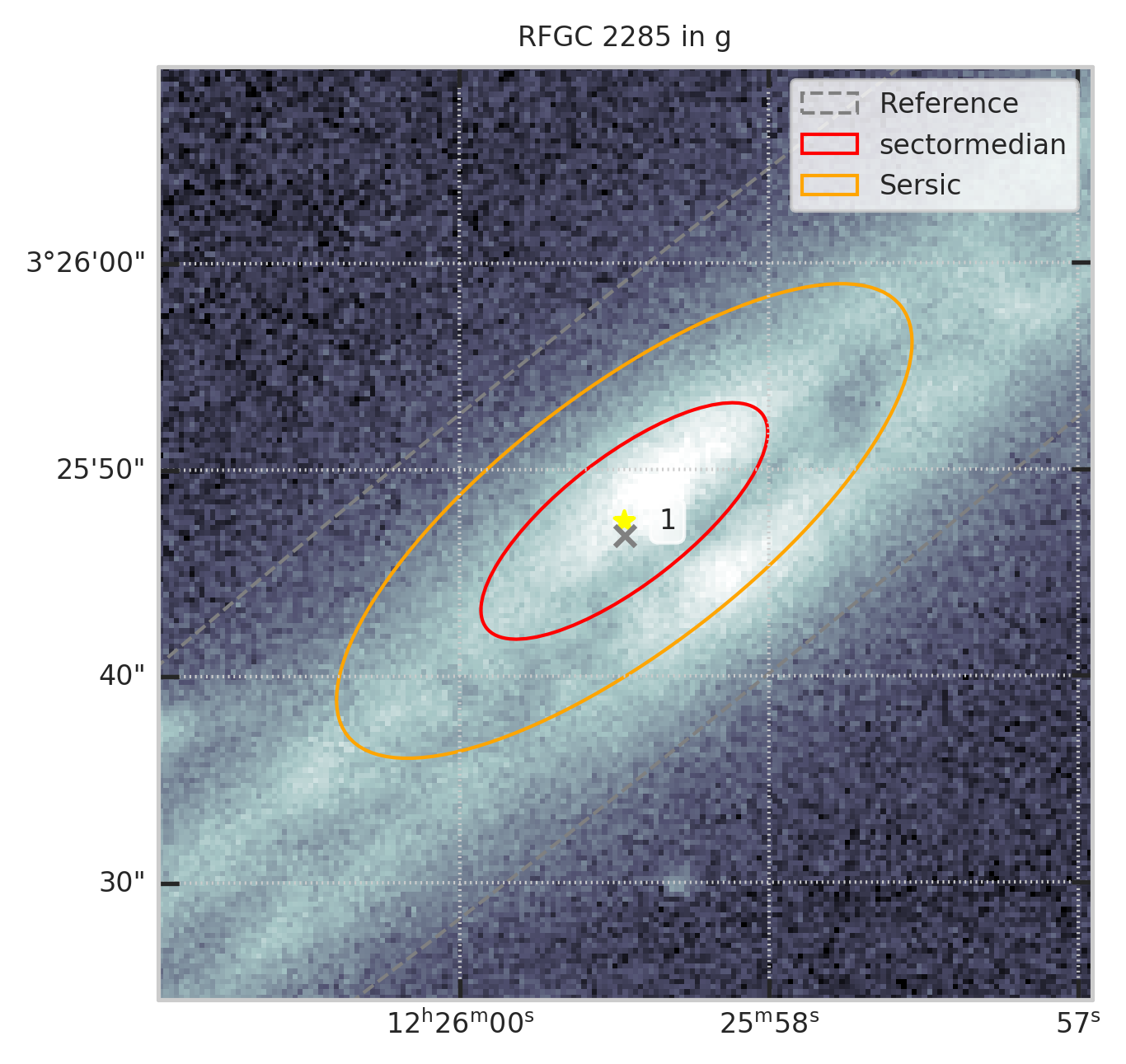
2. Оставим только те, которые есть в ForcedGalaxyShape в заданной полосе¶
filt = 'g'
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex"])
fig = plt.figure(figsize=(5,5))
t = wshape
subset = t[t.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=False, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
print(f"Осталось ~{len(t)/len(df)*100:.0f}% объектов")
ids_per_rfgc = t.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
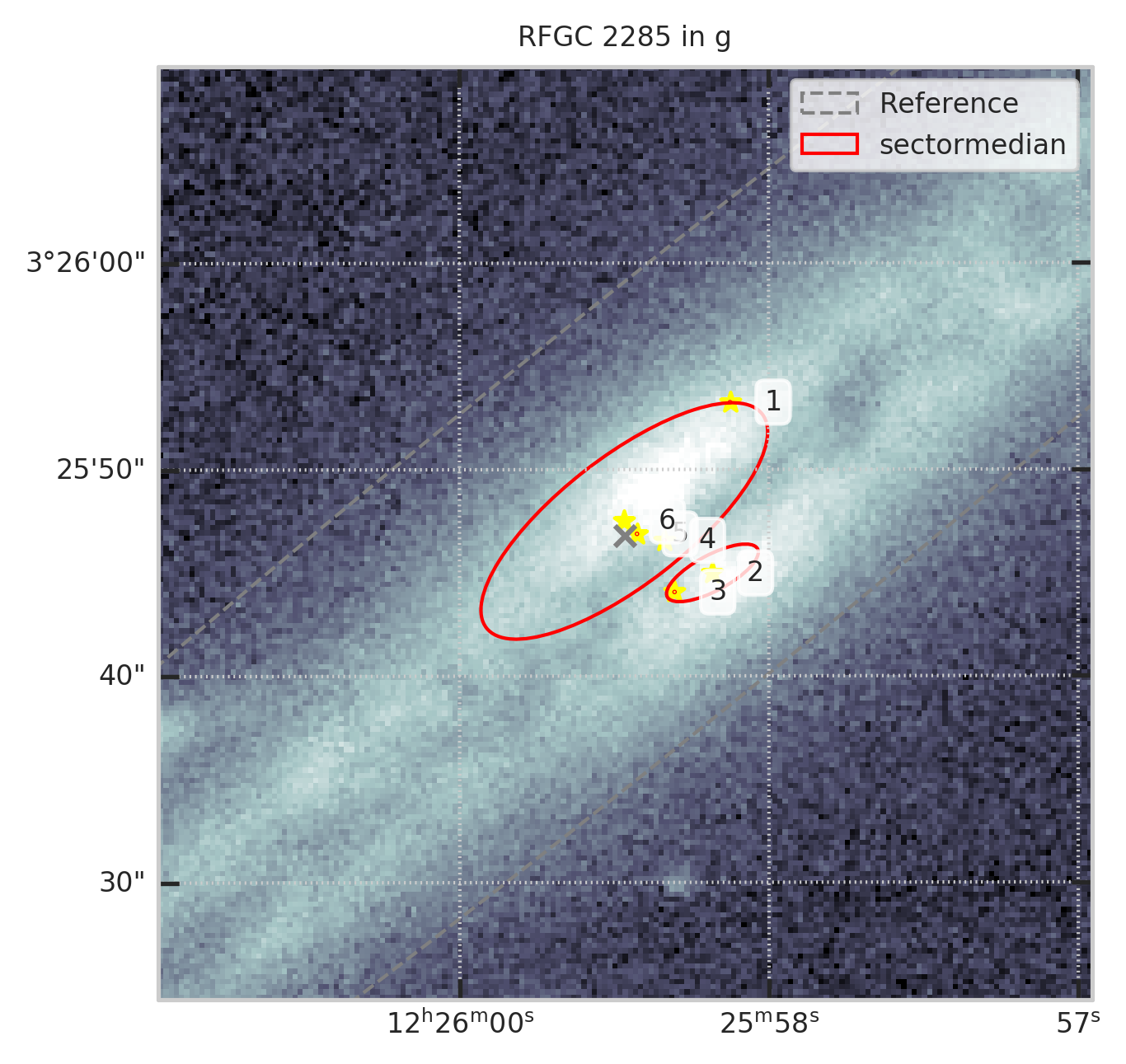
3. Оставим только те, для которых вписан Серсик в заданной полосе¶
filt = 'g'
fitted = wshape.dropna(subset=[f"{filt}SerRadius"])
fig = plt.figure(figsize=(5,5))
t = fitted
subset = t[t.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
print(f"Осталось ~{len(t)/len(df)*100:.1f}% объектов")
ids_per_rfgc = t.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
Все ещё остаются дубликаты
Посмотрим на объекты, которые PanSTARRS нашёл для галактики выше.
subset.loc[:, [f"{filt}SerRadius", f"{filt}SerAb", f"{filt}SerPhi", f"{filt}GalMajor", f"{filt}GalMinor", f"{filt}GalPhi"]]
Есть программный предел для радиуса Серсика: 0.05, позиционный угол и отношение осей при этом не определяются нормально
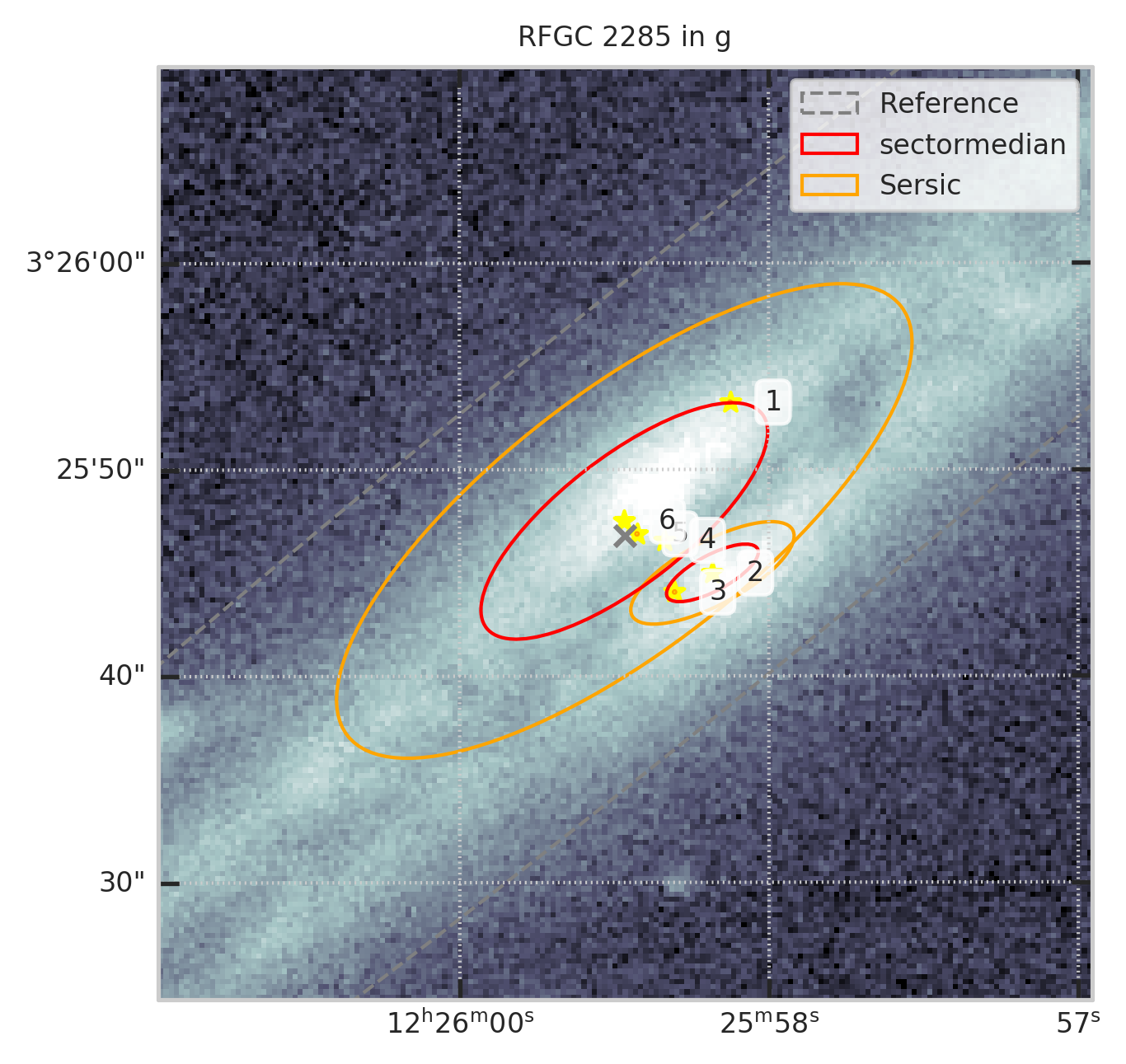
4. Отсеиваем программные ошибки вписания профиля¶
sane = fitted[fitted[f"{filt}SerA"] > 0.051]
fig = plt.figure(figsize=(5,5))
t = sane
subset = t[t.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
print(f"Осталось ~{len(t)/len(df)*100:.1f}% объектов")
ids_per_rfgc = t.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
Часть мусора ушла, остались протяженные источники, но мало что потерялось
5. Выбор согласованных по позиционному углу источников¶
aligned = sane[(np.abs(sane[f"{filt}GalPhi"] - sane["PA"]) < 10) &
(np.abs(sane[f"{filt}SerPhi"] - sane["PA"]) < 10)]
#aligned.index= range(1, len(aligned)+1)
fig = plt.figure(figsize=(5,5))
t = aligned
subset = t[t.RFGC == 2285]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=4, transform=LogStretch(10) + PercentileInterval(99.5))
print(f"Осталось ~{len(t)/len(df)*100:.1f}% объектов")
ids_per_rfgc = t.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
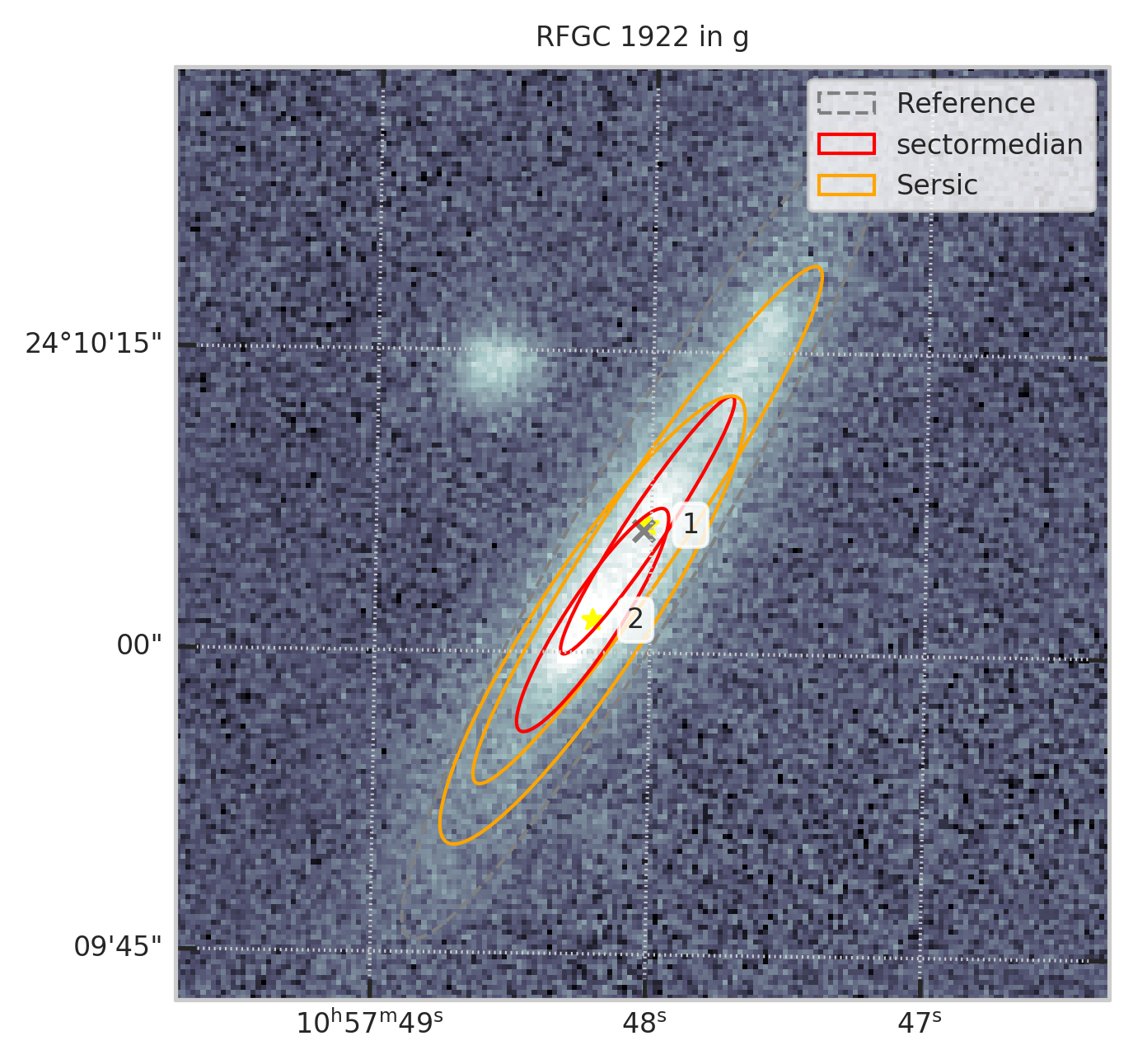
Выбор единственного наилучшего отождествления¶
Можно посмотреть, а в каких случаях объектов два
fig = plt.figure(figsize=(5,5))
t = aligned
subset = t[t.RFGC == 1922]
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=1, transform=LogStretch(10) + PercentileInterval(99.5))
subset.loc[:, [f"{filt}SerRadius", f"{filt}SerAb", f"{filt}SerPhi", f"{filt}SerMag",
f"{filt}GalMajor", f"{filt}GalMinor", f"{filt}GalPhi", f"{filt}GalMag"]]
Какие есть варианты оставить по одному отождествлению на галактику RFGC:
- Объект с наибольшей звездной величиной
- Объект с наибольшей большой полуосью
В первом случае выше вероятность отождествить центр галактики, видимо так более осмысленно
biggestsersic_aligned = aligned.sort_values(["RFGC",f"{filt}SerA"],ascending=[True, False]).groupby('RFGC').head(1)
biggestsermag_aligned = aligned.sort_values(["RFGC",f"{filt}SerMag"],ascending=[True, True]).groupby('RFGC').head(1)
summary = {}
masks_t = {}
sqlconds = {}
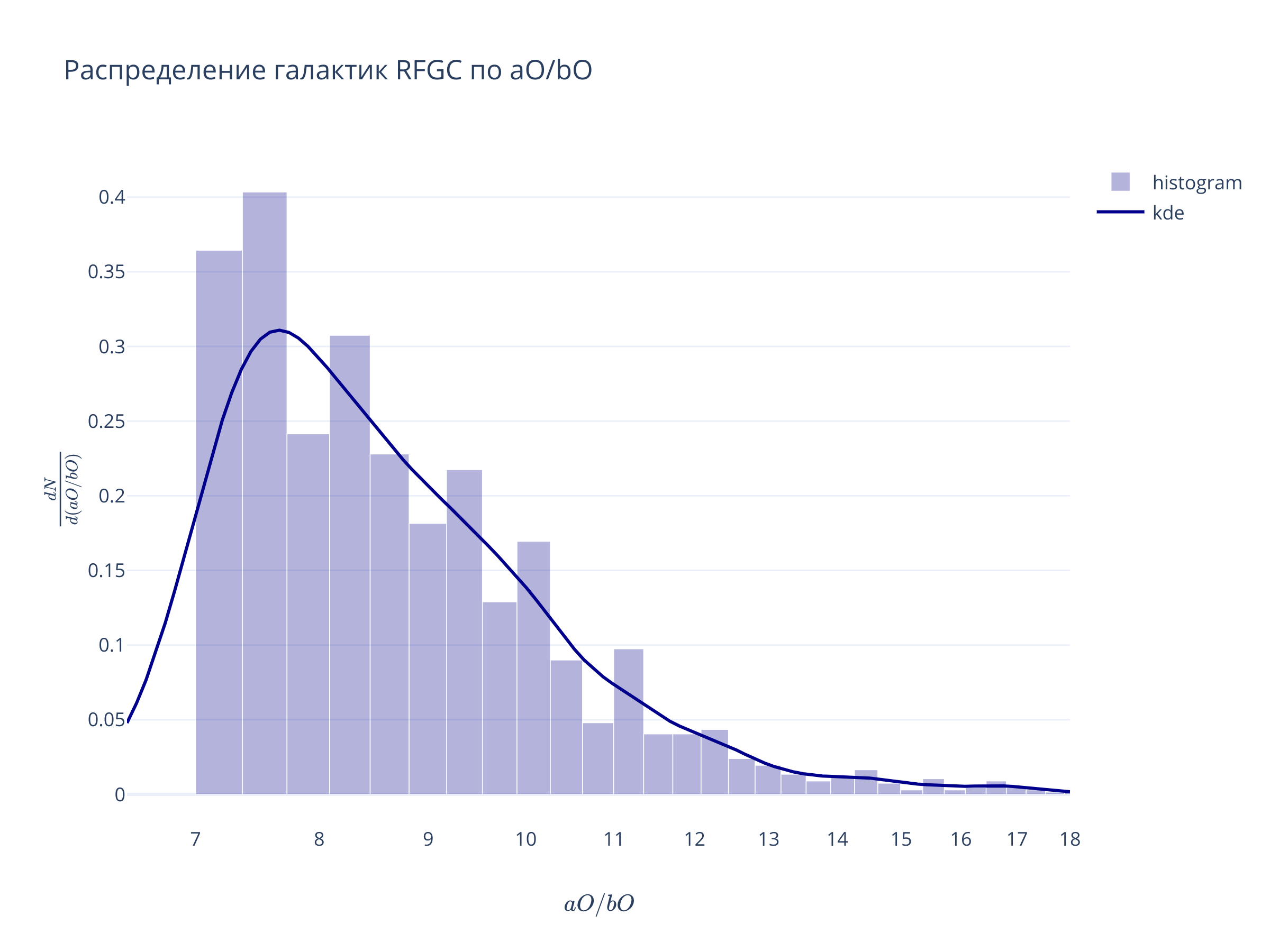
Подбор критериев отбора (g)¶
filt='g'
t = biggestsermag_aligned
t['BaO'] = 1/t['AbO']
from scipy.stats import gaussian_kde
yy, xx_e = np.histogram(t['BaO'], bins='auto', density=True)
xx = (xx_e[1:] + xx_e[:-1])/2
x1, x2 = np.log10(6.5), np.log10(18)
kernel = gaussian_kde(t['BaO'])
x_grid = np.logspace(x1, x2, 100)
fig = go.Figure(
data = [
go.Bar(x = xx, y = yy, width=xx[1]-xx[0], opacity=0.3, marker = {'color':'darkblue'}, name="histogram"),
go.Scatter(x = x_grid, y = kernel(x_grid), line = {'color':'darkblue'}, name = 'kde')
],
layout = go.Layout(xaxis = {'range':(x1, x2), 'type':'log', 'title':r"$aO/bO$"},
yaxis = {'title':r"$\frac{dN}{d (aO/bO)}$"},
title="Распределение галактик RFGC по aO/bO")
)
fig.show()
from code.plotutils import scatter_density_plotly
def scadensrfgc_ply(t, x, y, xrange, yrange, **kw):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
mode = kw.pop('mode', 'kde')
contours = kw.pop('contours', True)
logx = kw.pop('logx', False)
logy = kw.pop('logy', True)
plots_kw = dict(mode=mode, modepars={'bins':(30,30)}, alpha=0.9,
pointlabels=rfgc_names, contours=contours, **kw)
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=logy, logx=logx,
**plots_kw)
return fig
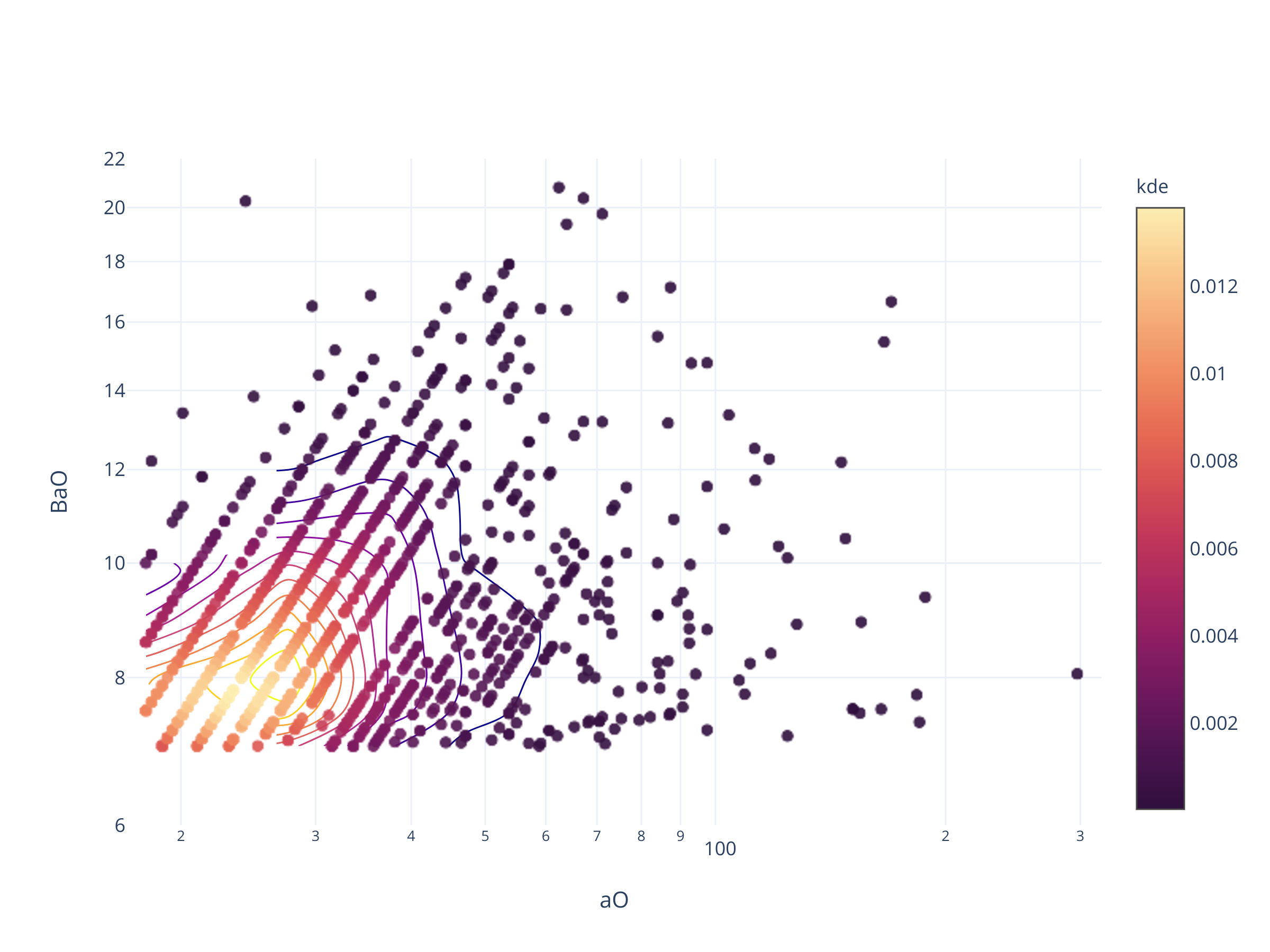
fig = scadensrfgc_ply(
biggestsermag_aligned,
'aO', 'BaO',
(17, 320), (6, 22),
logx=True
)
fig.update_layout(autosize=False, width=600, height=500)
fig
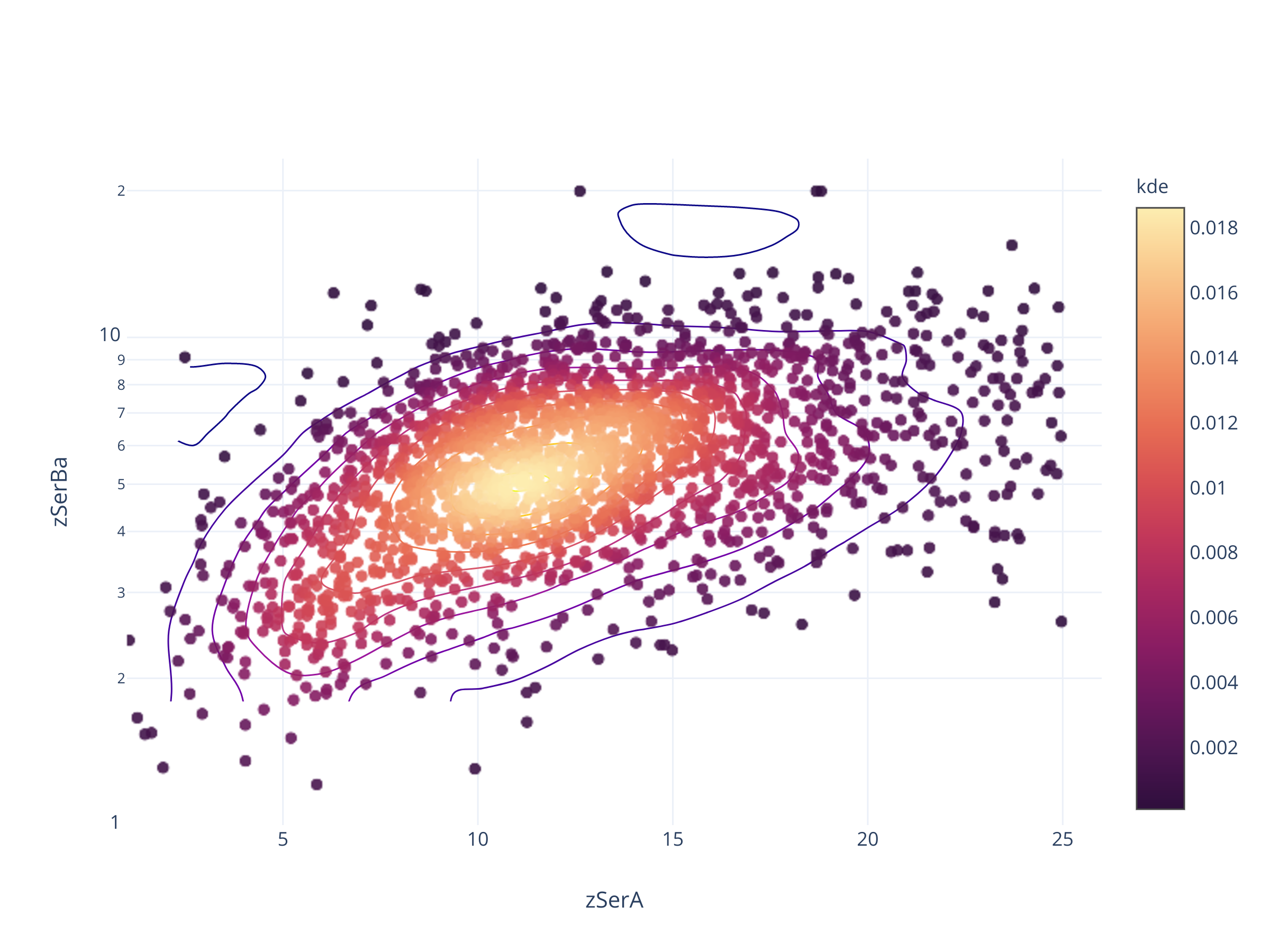
Посмотрим как оно может выглядеть для объектов PanSTARRS
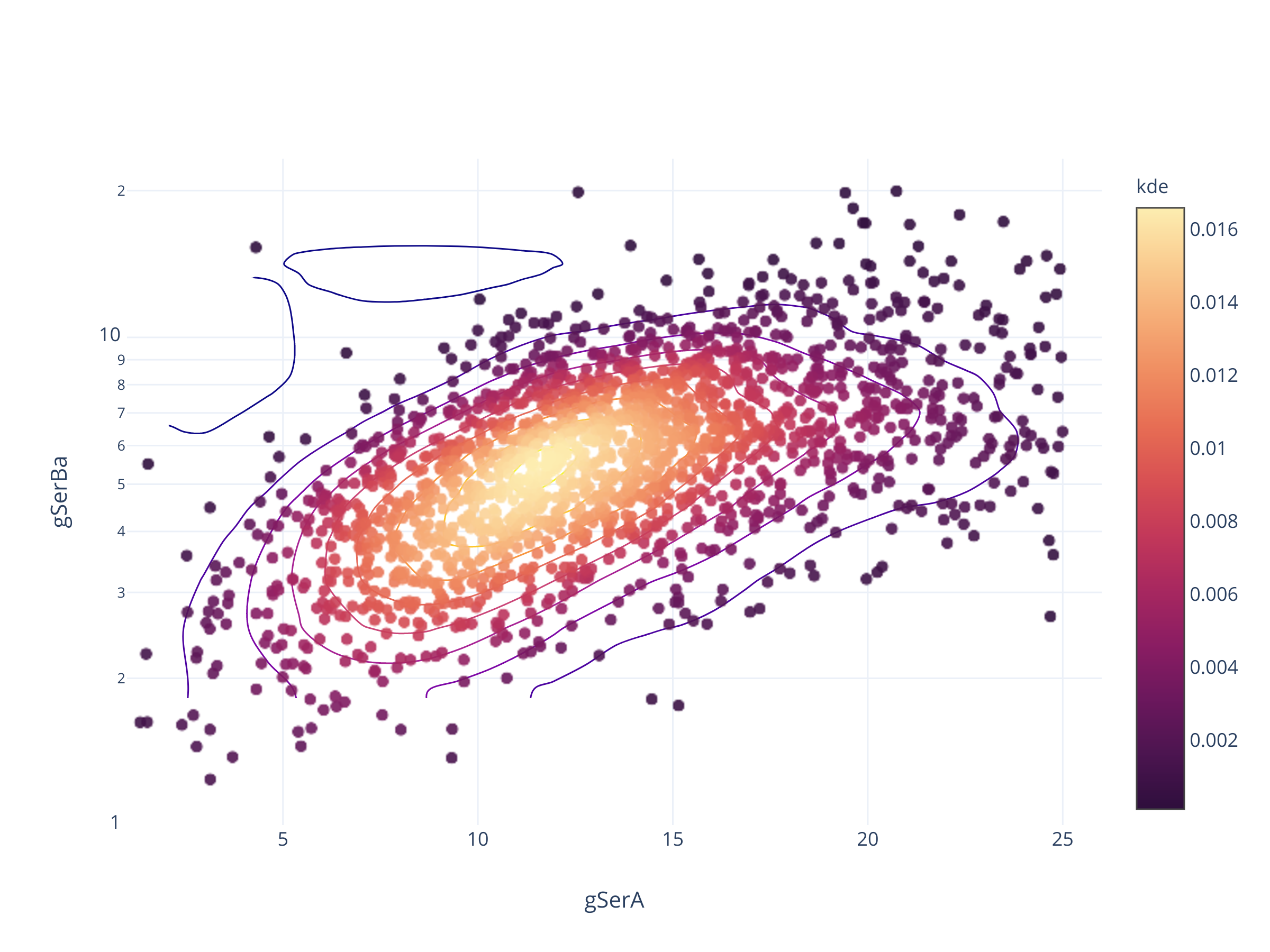
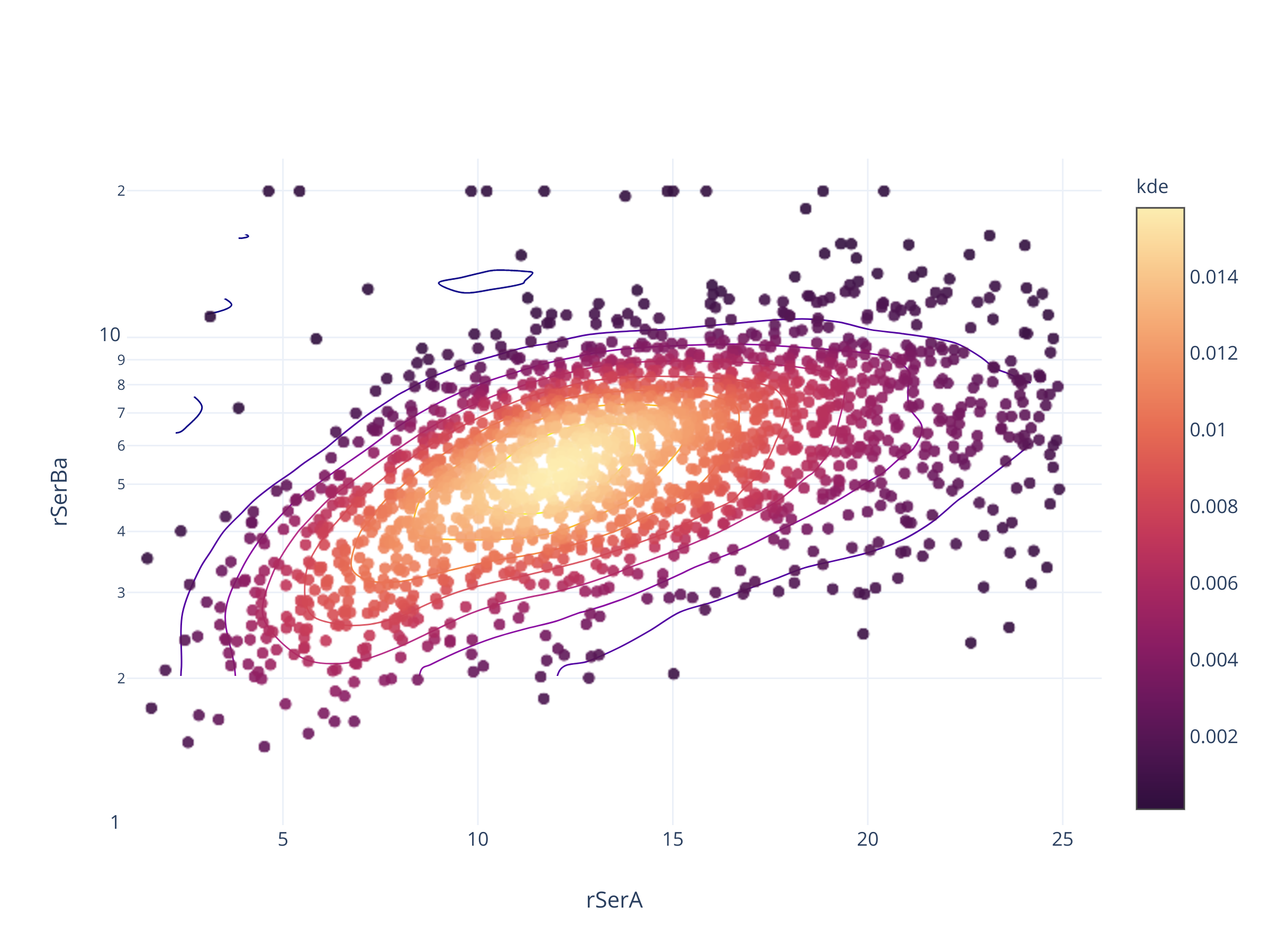
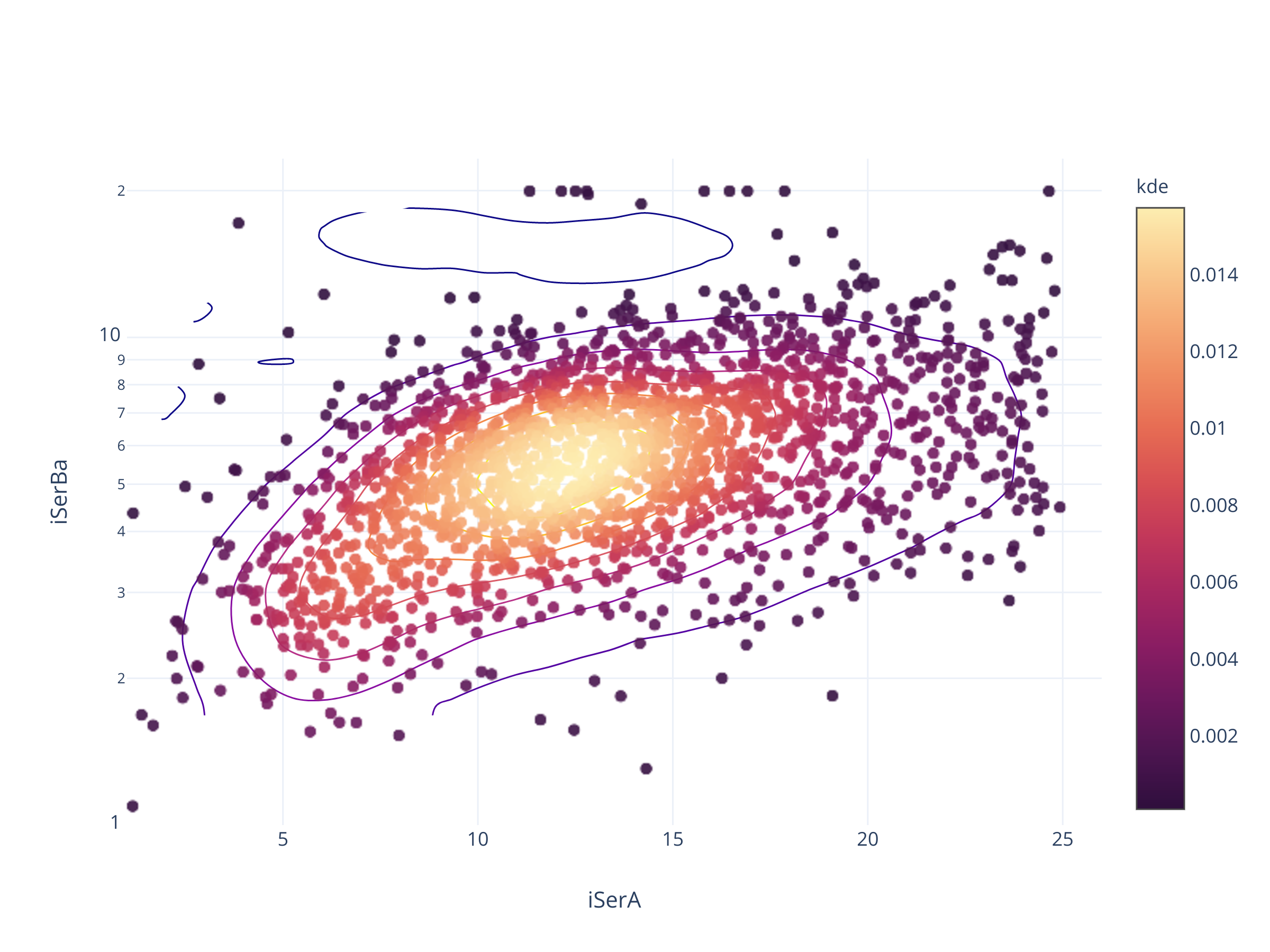
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 1/0.043)
)
fig.update_layout(autosize=False, width=600, height=500)
fig
Совсем не так.
В PanSTARRS, судя по всему, есть программный предел в 0.05 и 25.
Отрезать по $a/b > 7$ и $a > 18''$ бессмысленно -- ничего не останется, надо думать
0. Границы по большой полуоси¶
FWHM у PSF в обзоре $1''-2''$ (Глянул у пары галактик, но надо ещё уточнить)
Если у галактики полуось < $7''$, она будет видна с отношением осей $<7$ (а если нет, то это ошибка алгоритма). Получается, что надо отсекать где-то по $5'' - 7''$
t = biggestsermag_aligned
sizemask = {
5:(t[f'{filt}SerA'] > 5),
7:(t[f'{filt}SerA'] > 7),
}
summary[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index = t.index)}
sqlconds[filt] = {'all':f"{filt}SerRadius > 0.051"}
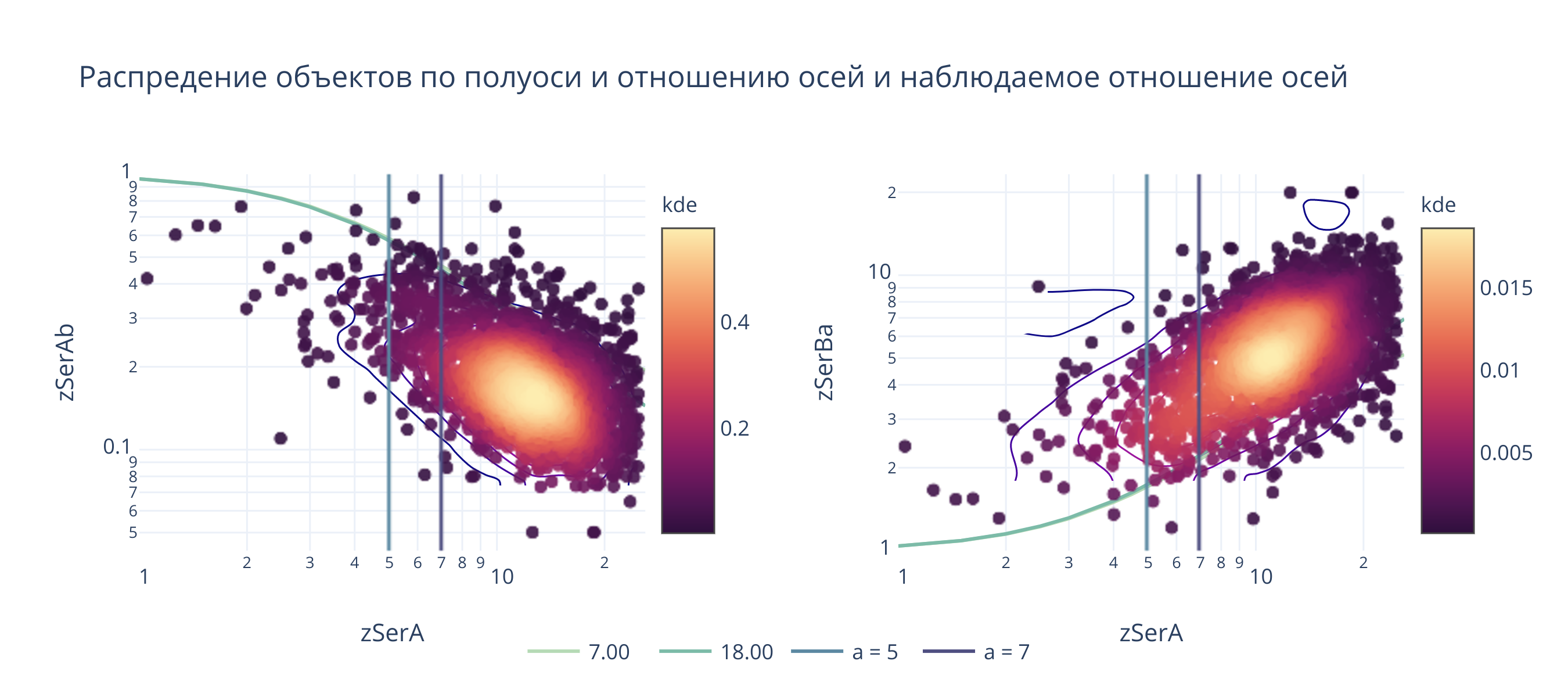
1. По «наблюдаемому» соотношению осей¶
Можно попробовать подобрать границу «круглости» с помощью этой функции
from code.plotutils import scatter_density_plotly, seaborn_to_plotly
def a_b_obs(a, a_b, sigma=1, inverse=False, ):
if inverse:
b = a * (1/a_b)
ratio = np.sqrt(b**2 + sigma**2) / np.sqrt(a**2 + sigma**2)
else:
b = a / a_b
ratio = np.sqrt(a**2 + sigma**2)/np.sqrt(b**2 + sigma**2)
return ratio
def b_a_obs(a, b_a, sigma=1):
return a_b_obs(a, 1/b_a, sigma, inverse=True)
def _():
x, y = f'{filt}SerA', f'{filt}SerAb'
y2 = f'{filt}SerBa'
xrange, yrange = (1, 26), (0.043, 1)
t = biggestsermag_aligned
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
sizes = np.linspace(*xrange, 51)
a_b_list = np.array([7, 25])
b_a_list = 1/a_b_list
N = len(a_b_list)
plots_kw = dict(mode="kde", modepars={'bins':(30,30)}, contours=True, alpha=0.9,
pointlabels=rfgc_names)
subplotlayout = dict(rows=1, cols=2)
fig = make_subplots(**subplotlayout)
fig.layout.update(
xaxis = {'domain':[0, 0.4]},
xaxis2 = {'domain':[0.6, 1]},
)
fig.layout.xaxis.type = 'log'
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=True, logx=True,
subplotpos={'row':1, 'col':1}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig = scatter_density_plotly(xx, 1/yy, xrange, (1/yrange[1], 1/yrange[0]), x, y2, logy=True, logx=True,
subplotpos={'row':1, 'col':2}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig.update_traces(
marker = {'colorbar':{'x':0.4}},
selector = {'type':'scattergl'},
row = 1, col = 1)
fig.update_traces(
marker = {'colorbar':{'x':1}},
selector = {'type':'scattergl'},
row = 1, col = 2)
fig.update_layout(showlegend=True, legend_orientation="h")
pallete = sb.cubehelix_palette(4, start=.5, gamma=0.8, rot=-0.7, dark=0.25, light=0.75)
ply_chx = seaborn_to_plotly(pallete)
nncurves = len(fig.data)
for i, a_b in enumerate(a_b_list):
fig.add_trace(
go.Scattergl(
x = sizes,
y = b_a_obs(sizes, 1/a_b),
name="%.2f" % a_b, showlegend=True,
legendgroup="%.2f" % a_b,
hoverinfo='name',
line = {'color':ply_chx[i][1]}
), row=1, col=1)
fig.add_trace(
go.Scattergl(
x = sizes,
y = a_b_obs(sizes, a_b),
name = "%.2f" % a_b,
hoverinfo='name', showlegend=False,
line = {'color':ply_chx[i][1]}
), row=1, col=2)
for i, x in enumerate([5, 7]):
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo = "name", legendgroup=f"a = {x}"),
row=1, col=1)
yrange2 = (1/yrange[1], 1/yrange[0])
fig.add_trace(go.Scattergl(x = [x, x], y = yrange2, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo="name", showlegend=False),
row=1, col=2)
fig.layout.legend.x = 0.3
fig.layout.legend.y = -0.2
fig.layout.legend.tracegroupgap = 30
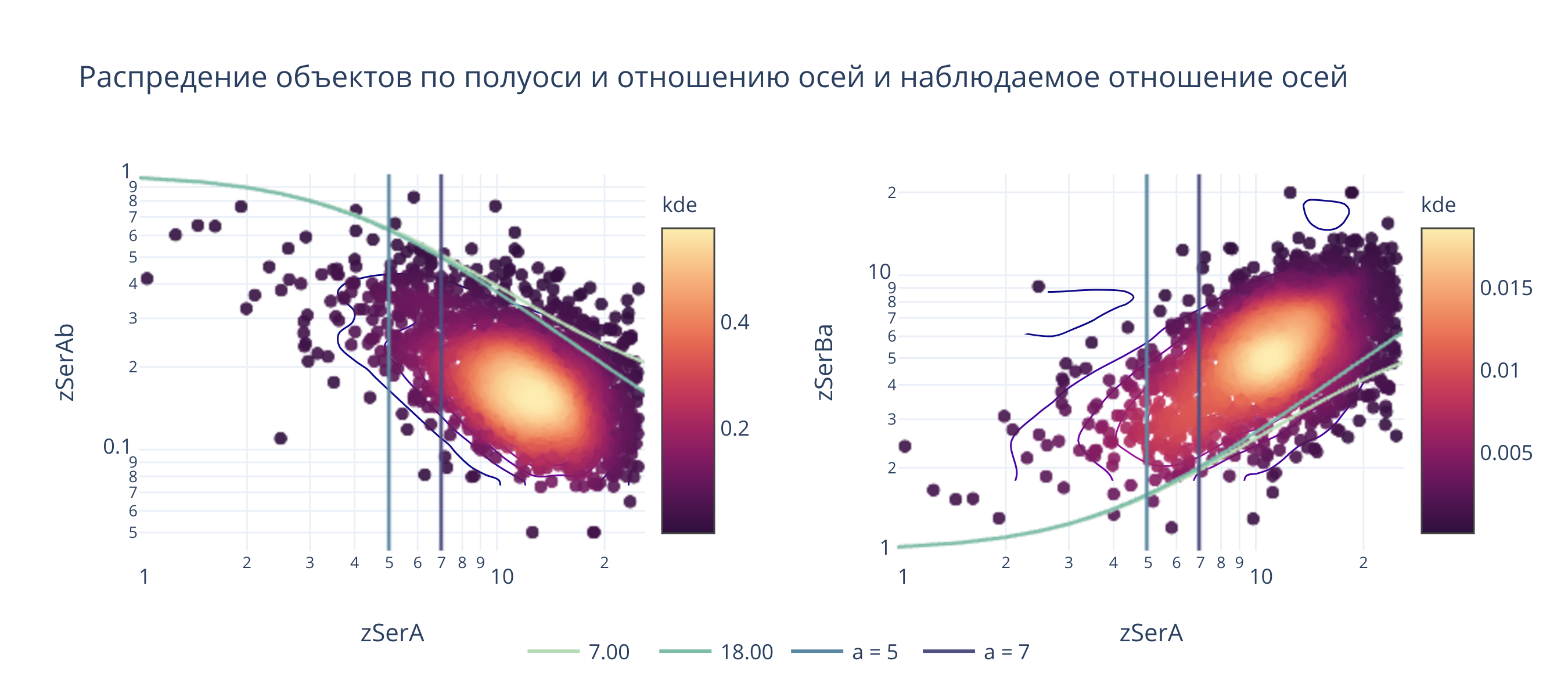
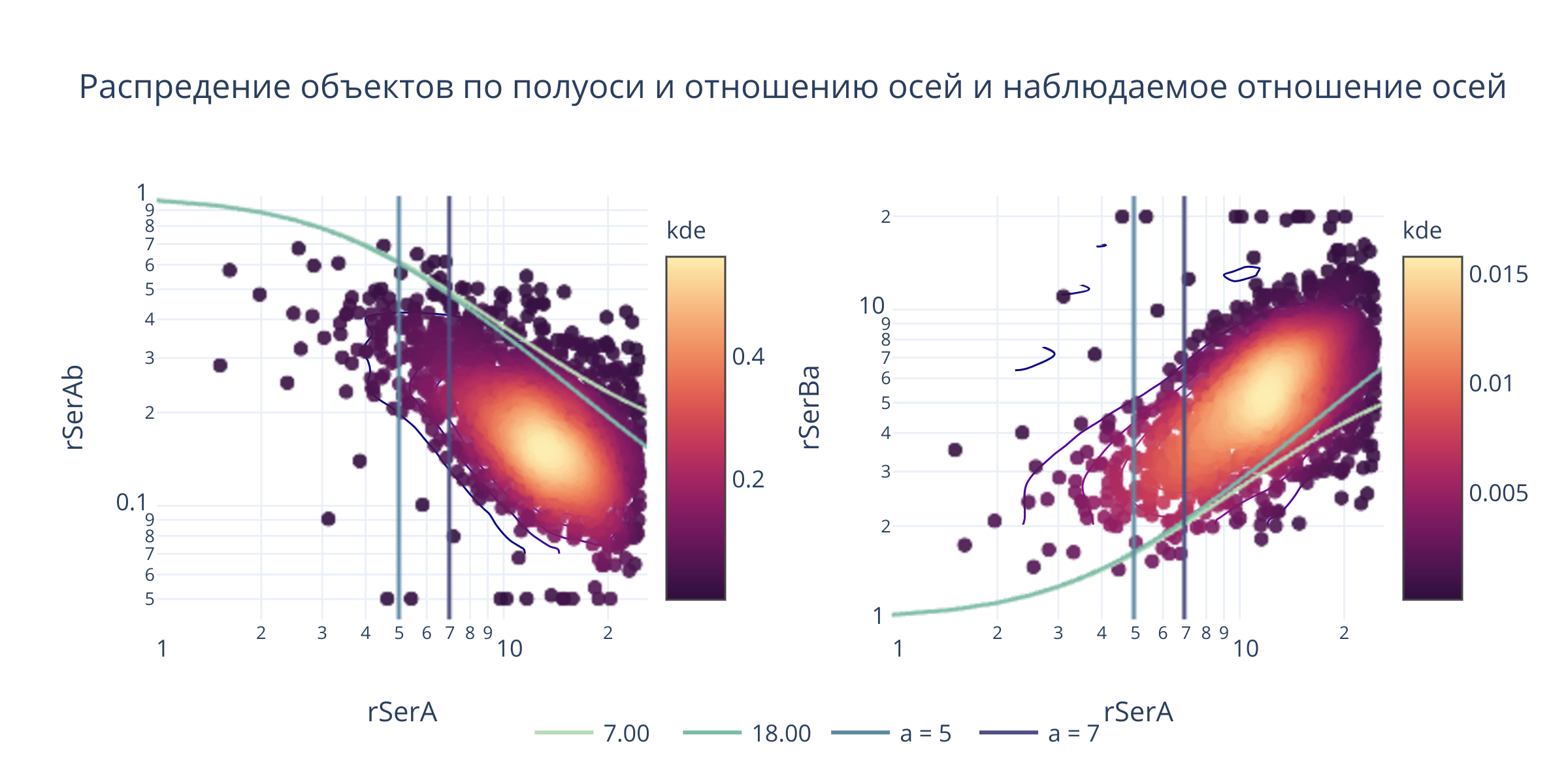
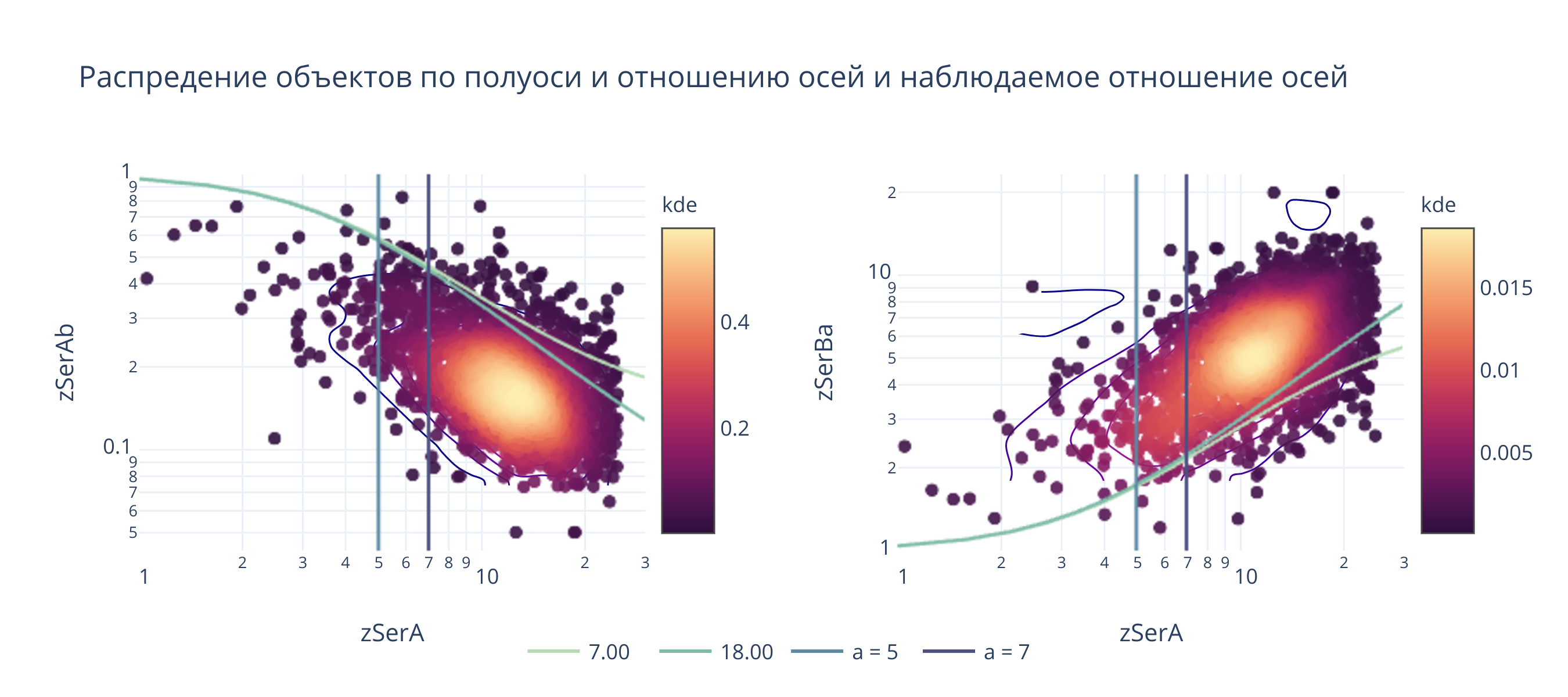
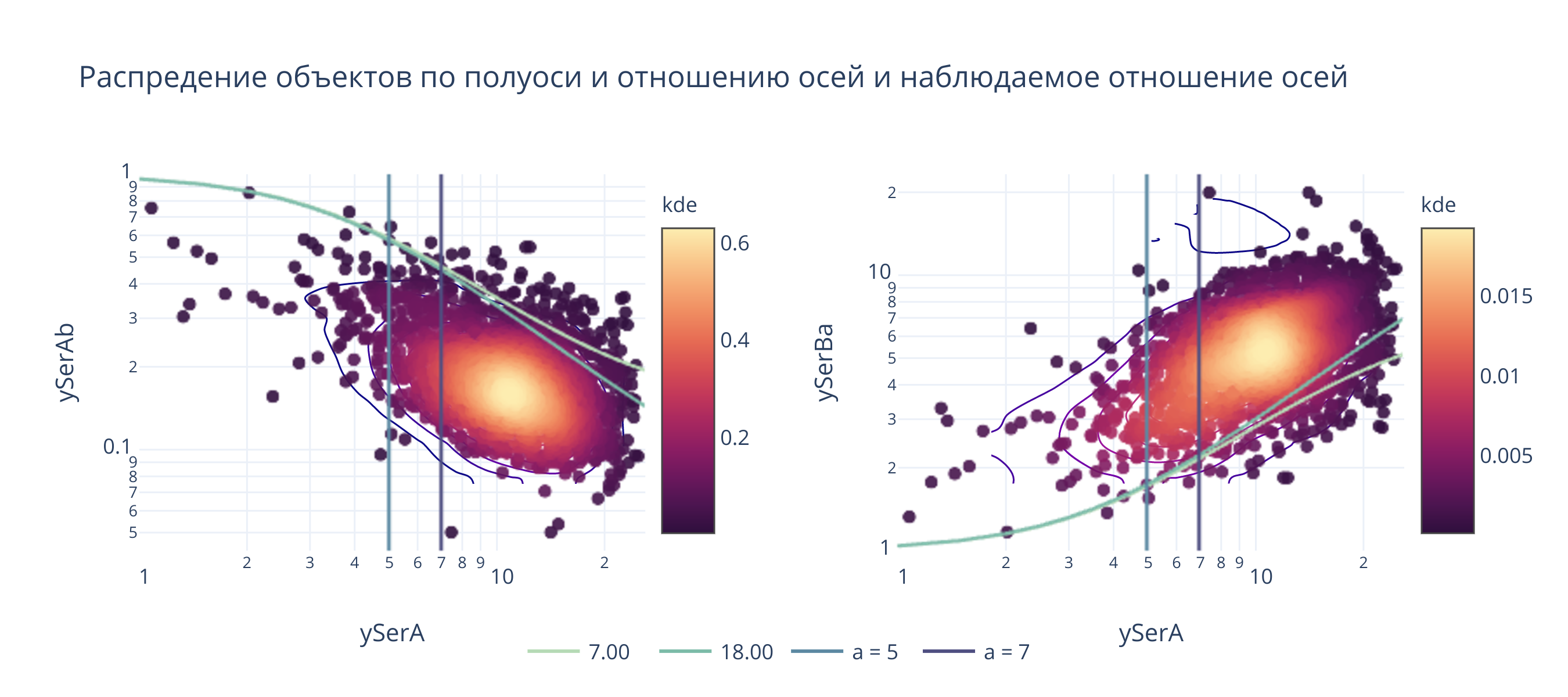
fig.layout.title = "Распредение объектов по полуоси и отношению осей и наблюдаемое отношение осей"
fw = go.FigureWidget(fig)
@interact(a_b1 = (1, 30, .1), a_b2 = (1, 30, .1), sigma=(1, 10, 0.1))
def update(a_b1 = 7, a_b2 = 18, sigma=4):
with fw.batch_update():
for i, (a_b, inverse) in enumerate([(a_b1, True), (a_b1, False), (a_b2, True), (a_b2, False)],
start=nncurves):
fw.data[i].update(
y = a_b_obs(sizes, a_b, sigma, inverse=inverse),
name = "%.2f" % a_b)
return fw
_().show(height = 400, width=900)
def sel_obs_ratio(t, r_obs, sigma, amin):
key = f'r_obs={r_obs:d} sigma={sigma:.2f} a>{amin:d}'
masks_t[filt][key] = (
(t[f'{filt}SerAb'] < a_b_obs(t[f'{filt}SerA'], r_obs, sigma=sigma, inverse=True)) &
sizemask[amin]
)
sqlconds[filt][key] = f"{filt}SerAb < SQRT(SQUARE({filt}SerRadius/{r_obs:.2f}) + {sigma**2:.2f}) / SQRT(SQUARE({filt}SerRadius) + {sigma**2:.2f}) AND {filt}SerRadius > {amin:.2f}"
sel_obs_ratio(biggestsermag_aligned, 7, 4, 5)
sel_obs_ratio(biggestsermag_aligned, 7, 4, 7)
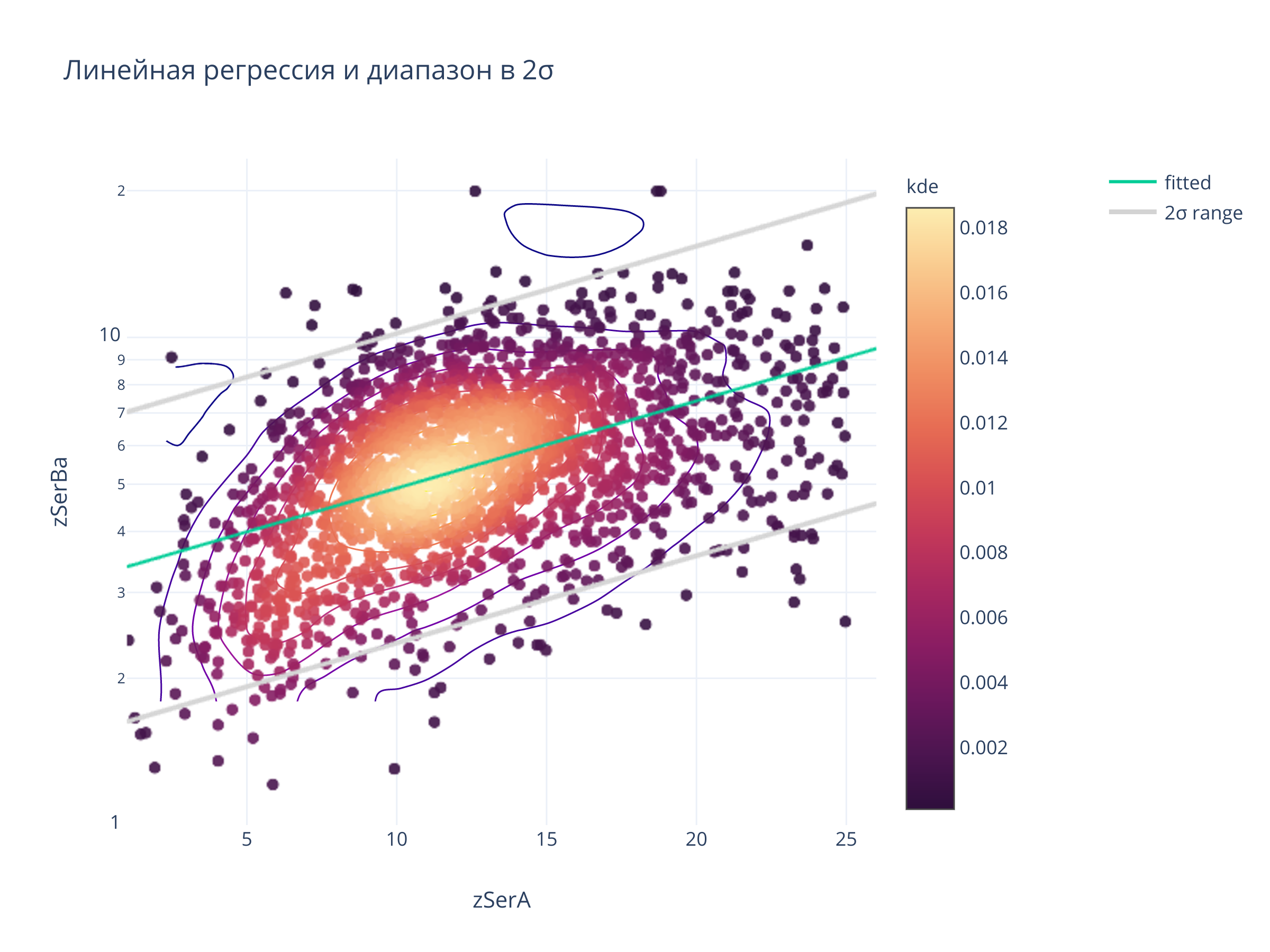
2. По дисперсии линейной регрессии¶
import statsmodels.api as sm
from astropy.modeling import models, fitting
def compute_linregr_pars(t, x, y, xrange, yrange, logx=False, logy=True):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
xx_reg = (t.loc[mask, x])
yy_reg = (t.loc[mask, y])
xx_reg = np.log10(xx_reg) if logx else xx_reg
yy_reg = np.log10(yy_reg) if logy else yy_reg
variables = pd.DataFrame({
'const':np.ones_like(xx_reg),
'x':xx_reg,
#'x2':xx**2,
})
linmodel = sm.OLS(yy_reg, variables)
a,b,c,d,e = [0]*5
mod = Namespace()
linresults = linmodel.fit()
mod.eps = linresults.resid
mod.b,mod.a = linresults.params
hist, xe = np.histogram(mod.eps)
xe = (xe[1:]+xe[:-1])/2
ini = models.Gaussian1D()
fitter = fitting.LevMarLSQFitter()
fit = fitter(ini, xe, hist)
mod.sigma_total = fit.stddev
return mod
xrange, yrange = (1, 26), (1, 23.26)
m = compute_linregr_pars(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
x_grid = np.linspace(*xrange, 100)
fitted = 10**(m.a*x_grid + m.b)
upperbnd = 10**(m.a*x_grid + m.b + 2*m.sigma_total)
lowerbnd = 10**(m.a*x_grid + m.b - 2*m.sigma_total)
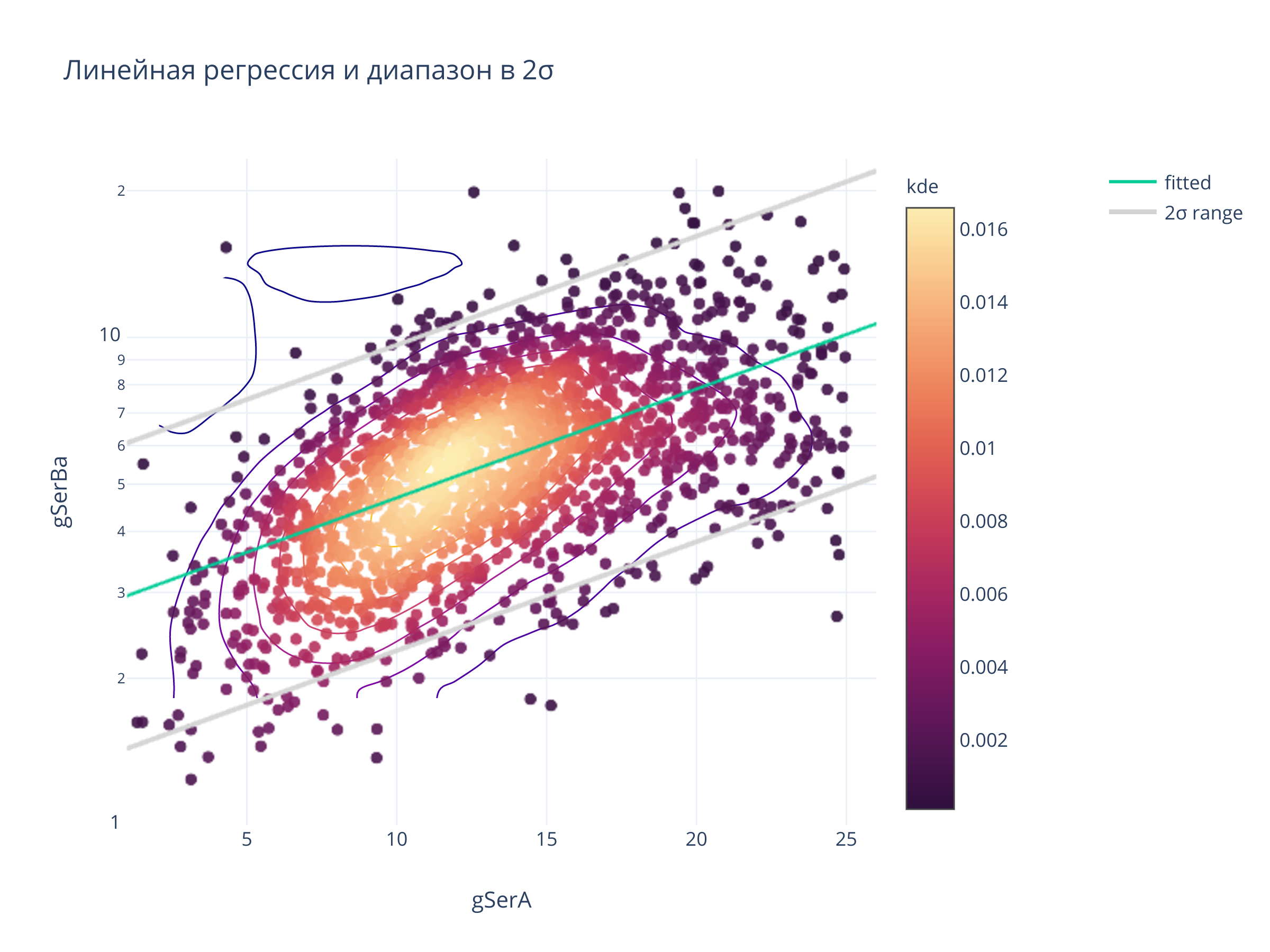
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
fig.add_trace(go.Scattergl(x = x_grid, y = fitted, name='fitted'))
fig.add_trace(go.Scattergl(x = np.hstack([x_grid, [x_grid[-1]+1,x_grid[-1]+1], np.flip(x_grid)]),
y = np.hstack([upperbnd, [upperbnd[-1], lowerbnd[-1]], np.flip(lowerbnd)]), line = {'color':'lightgray', 'width':3},
name='2σ range'))
fig.layout.legend.x = 1.3
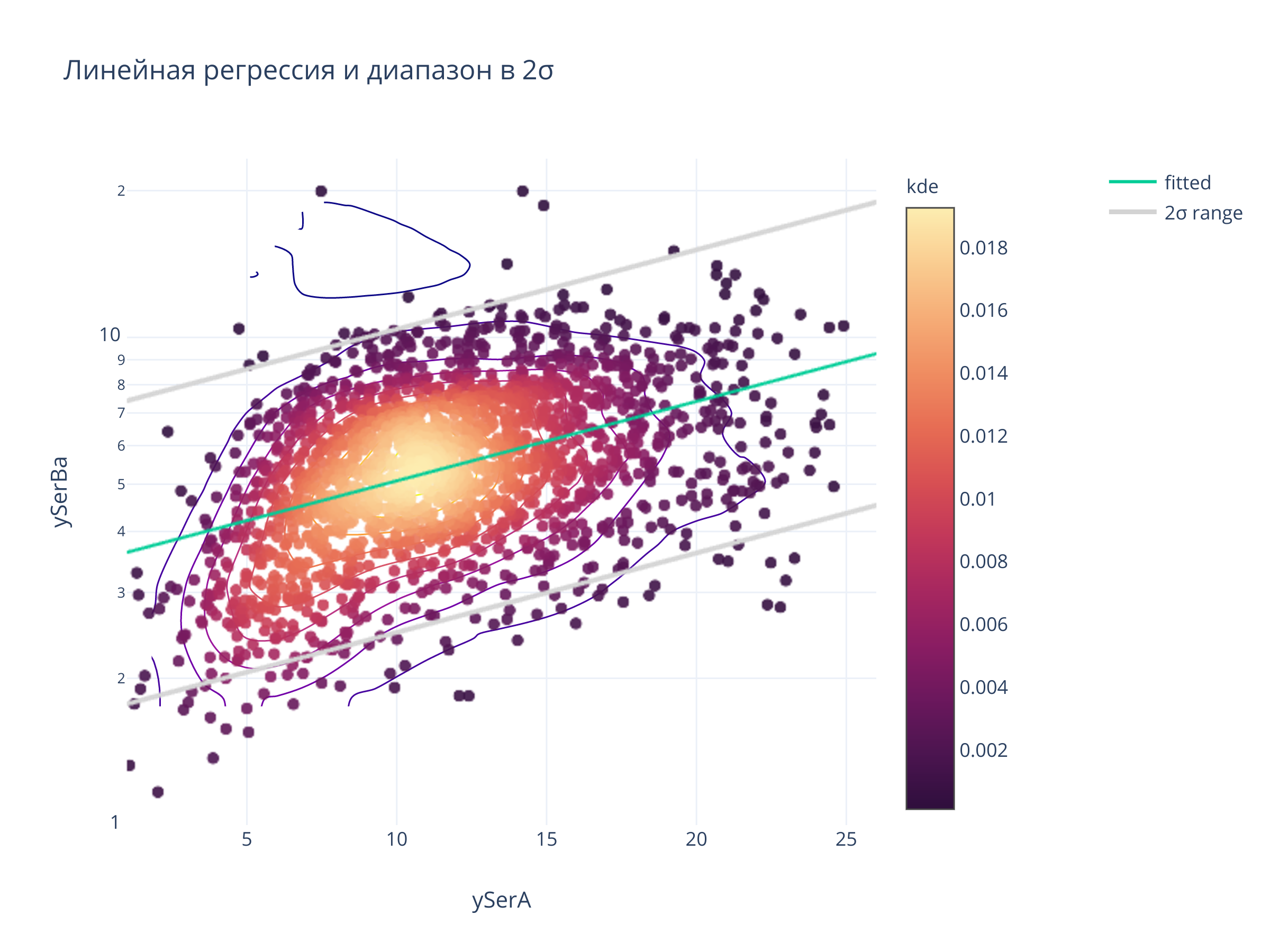
fig.update_layout(
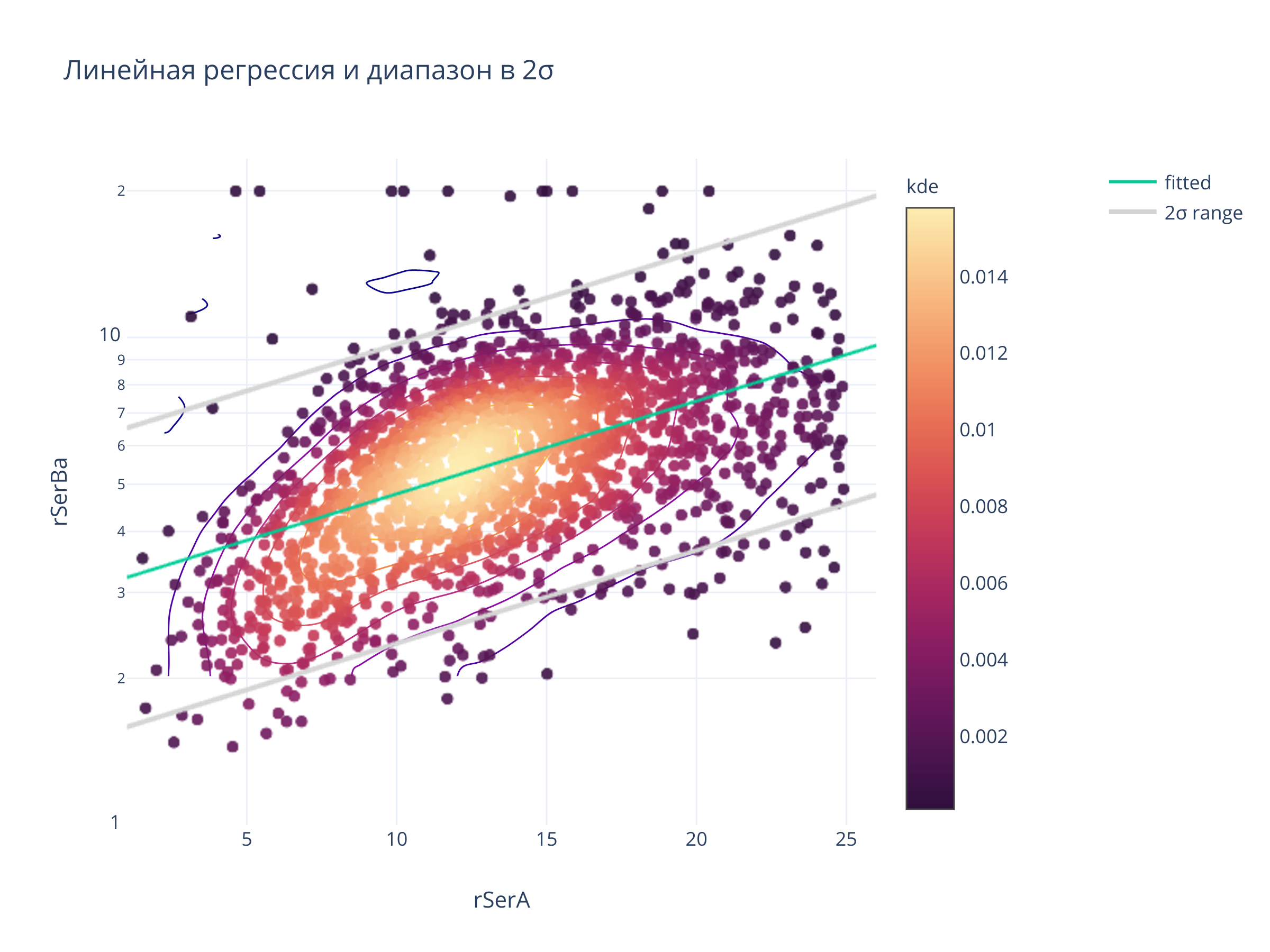
title = "Линейная регрессия и диапазон в 2σ",
clickmode = "event+select",
autosize=False,width=700, height=500)
fig.show()
def sel_lbnd_linreg(t, amin):
key = f'2 sigma a>{amin:d}'
masks_t[filt][key] = (
(np.log10(t[f'{filt}SerBa']) > m.a*t[f'{filt}SerA']+m.b - 2*m.sigma_total) &
(t[f'{filt}SerA'] > amin)
)
sqlconds[filt][key] = f"-LOG10({filt}SerAb) > {m.a:.5f}*gSerRadius + {m.b:.5f} - {2*m.sigma_total:.5f} AND {filt}SerRadius > {amin:.2f}"
#t = biggestsermag_aligned
sel_lbnd_linreg(biggestsermag_aligned, 5)
sel_lbnd_linreg(biggestsermag_aligned, 7)
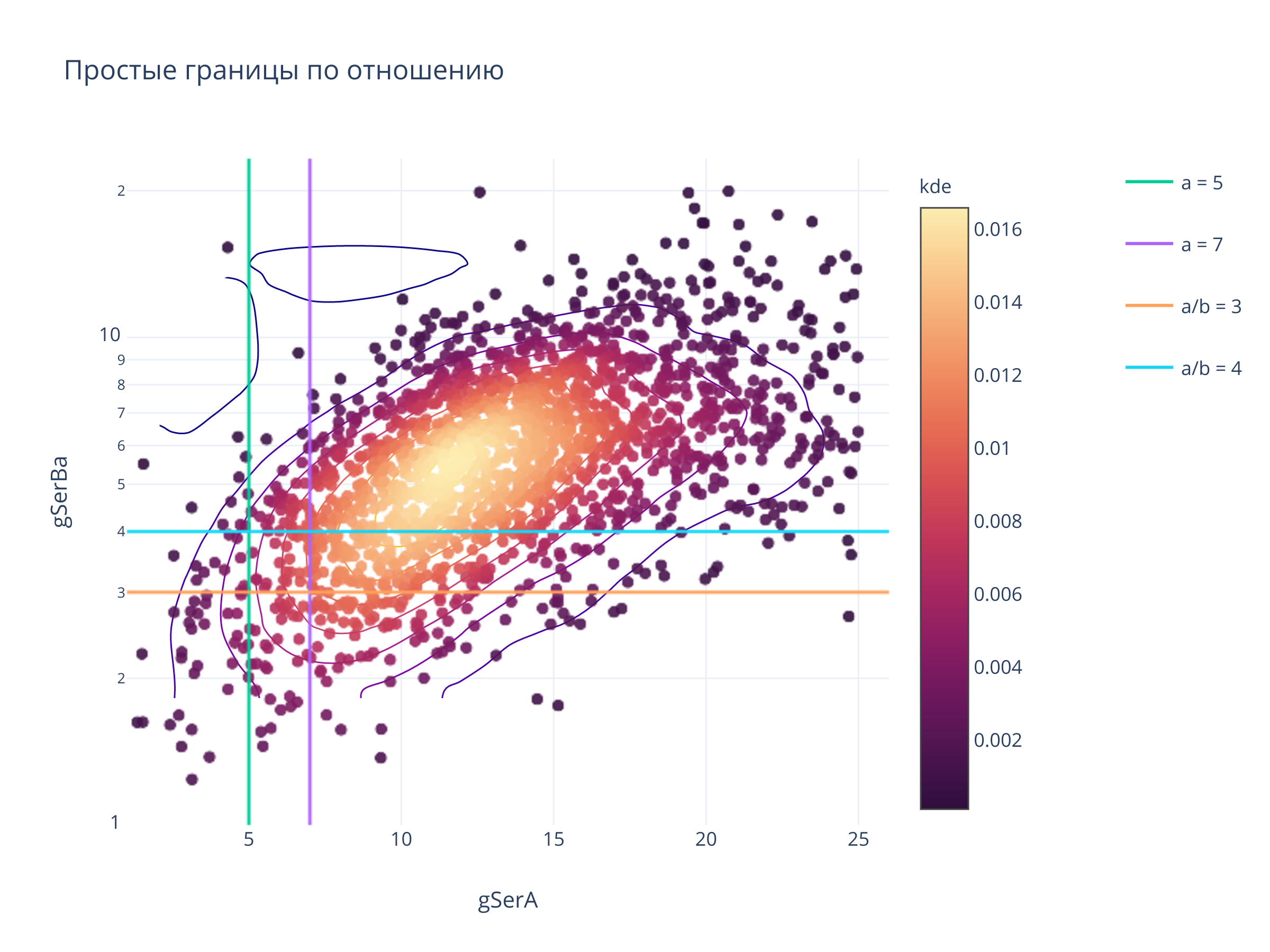
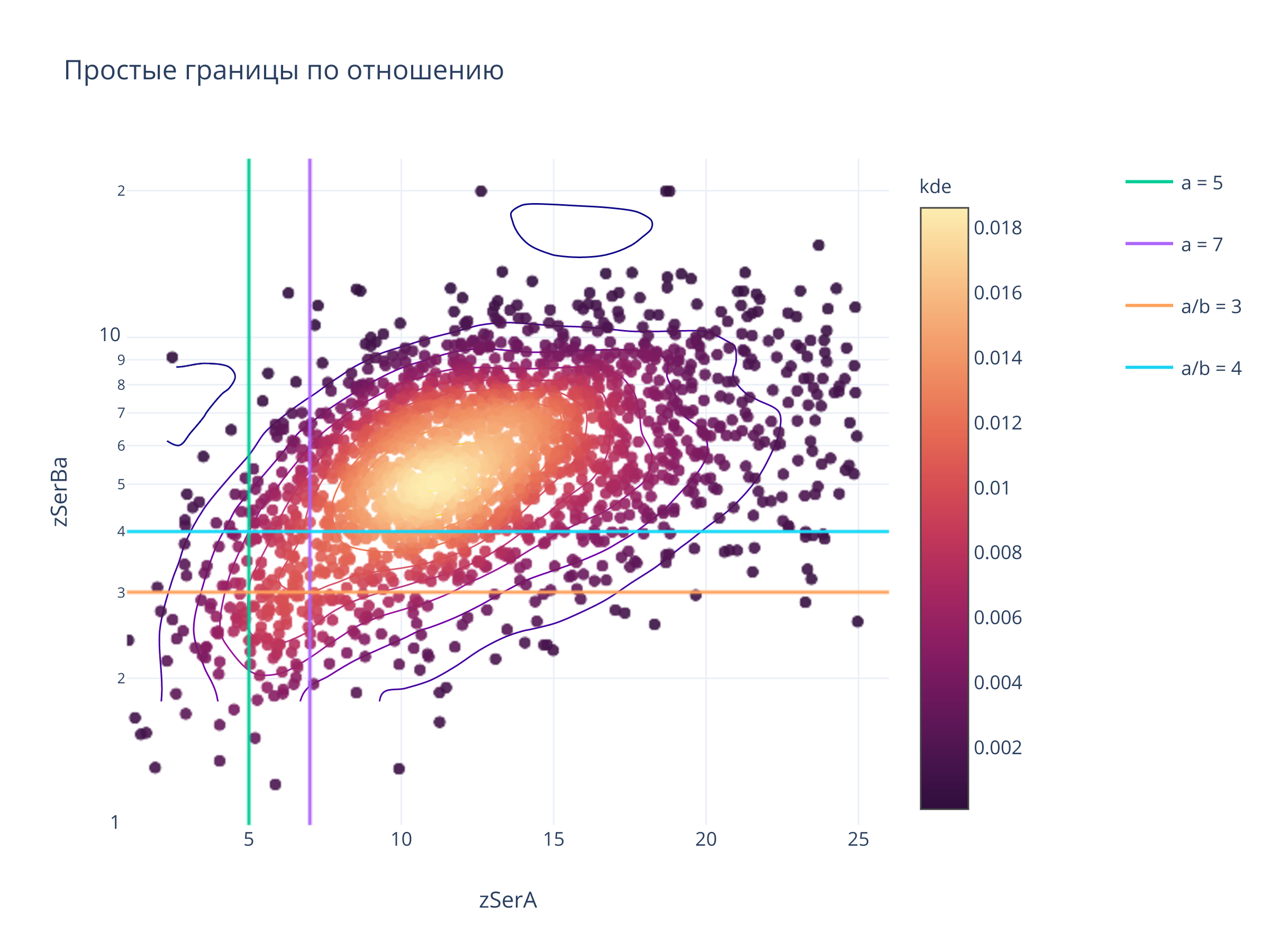
3. Простые границы¶
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 23.26)
)
x_grid = np.linspace(*xrange, 100)
fig.layout.legend.x = 1.3
fig.layout.legend.tracegroupgap = 20
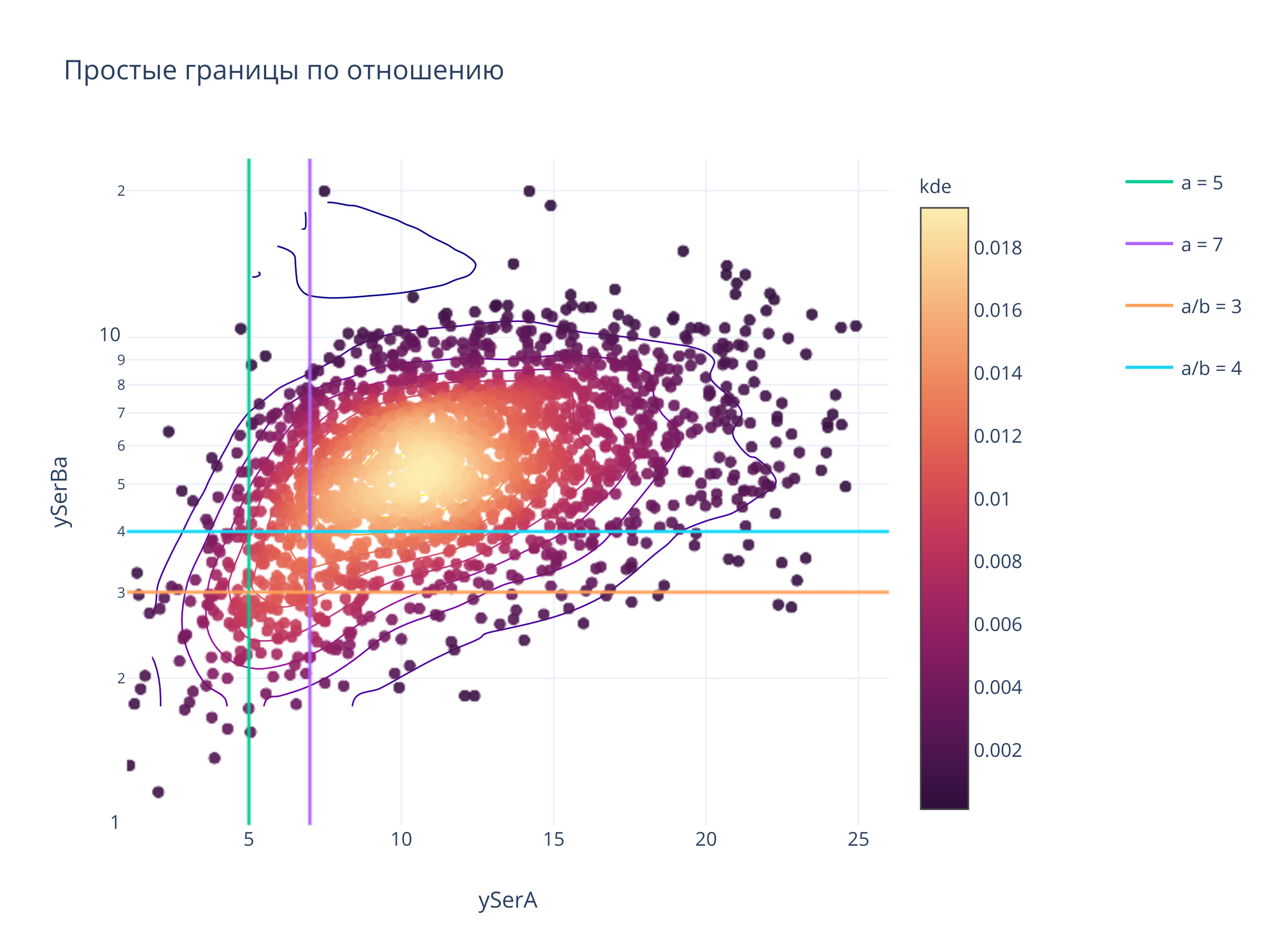
fig.update_layout(
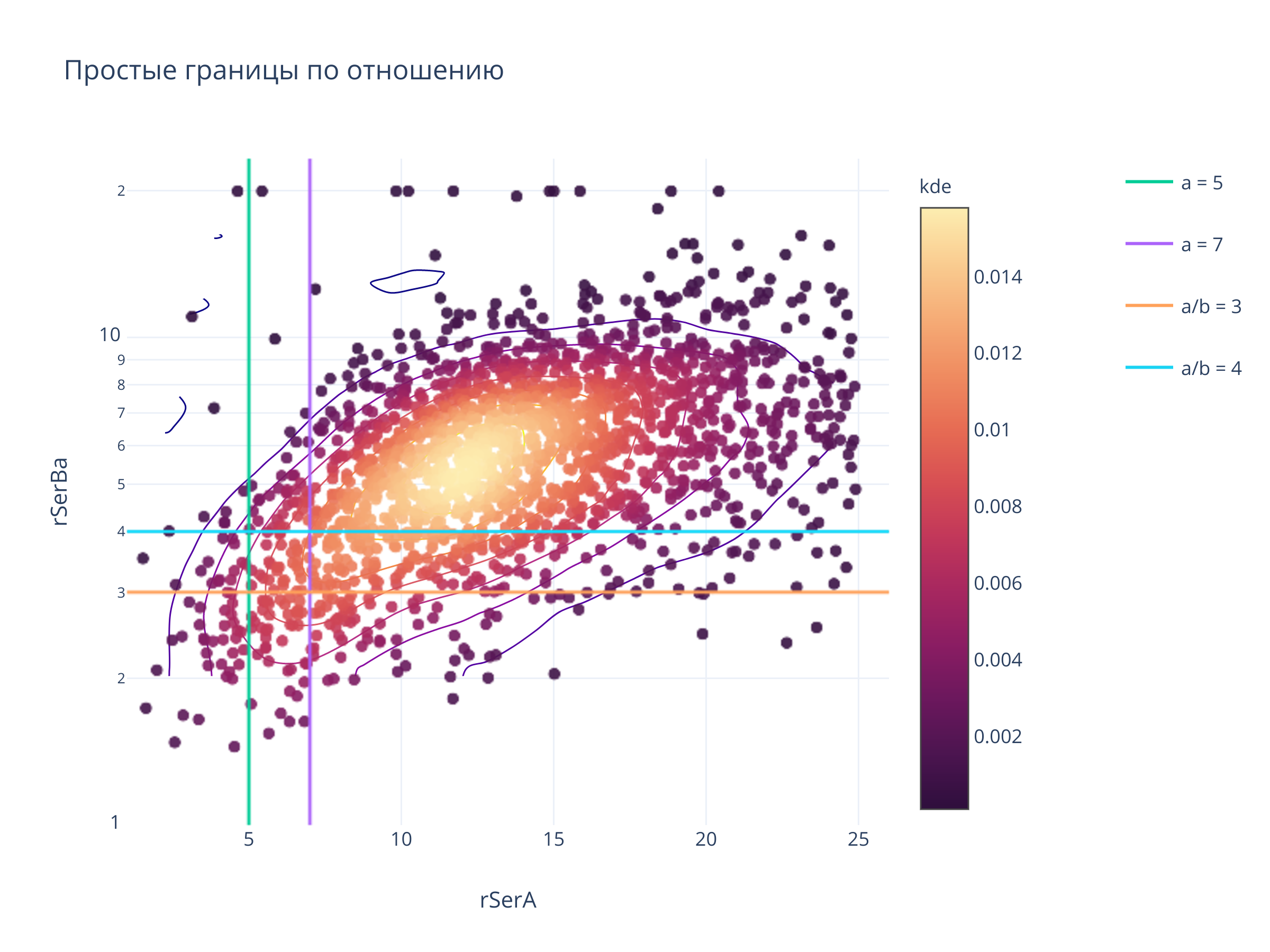
title = "Простые границы по отношению",
clickmode = "event+select",
autosize=False,width=800, height=600)
for x in [5, 7]:
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", name=f"a = {x}", legendgroup=f"a = {x}"))
for y in [3, 4]:
fig.add_trace(go.Scattergl(x = xrange, y = [y,y],mode = "lines",name=f"a/b = {y}", legendgroup = f"a/b = {y}"))
fig
def sel_tresh(t, ratio, amin):
key = f'a/b>{ratio:d} a>{amin:d}'
masks_t[filt][key] = (t[f'{filt}SerA'] > amin) & (t[f'{filt}SerBa'] > ratio)
sqlconds[filt][key] = f"{filt}SerRadius > {amin:.2f} AND {filt}SerAb < {1/ratio:.5f}"
sel_tresh(biggestsermag_aligned, 3, 5)
sel_tresh(biggestsermag_aligned, 3, 7)
sel_tresh(biggestsermag_aligned, 4, 5)
sel_tresh(biggestsermag_aligned, 4, 7)
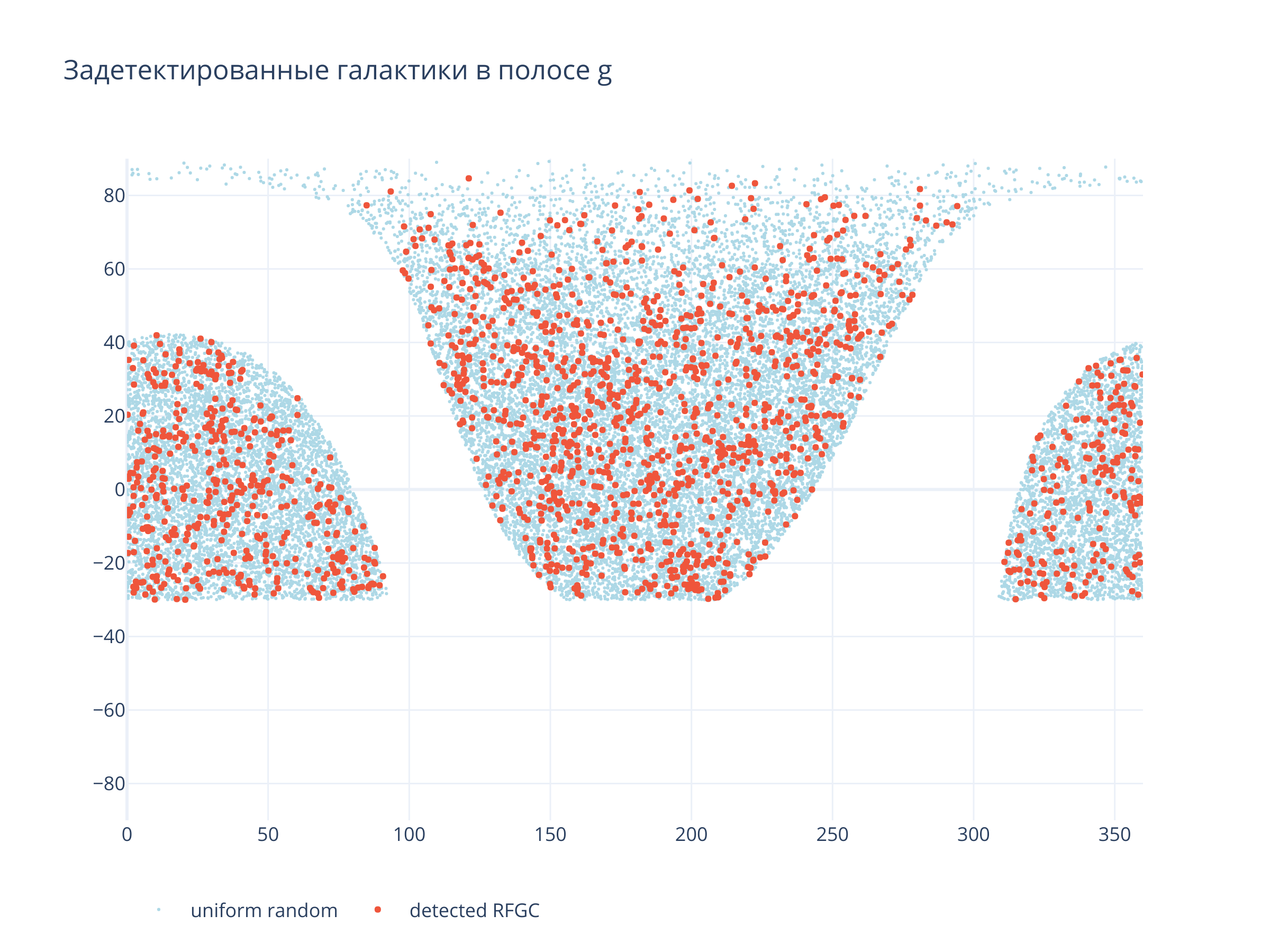
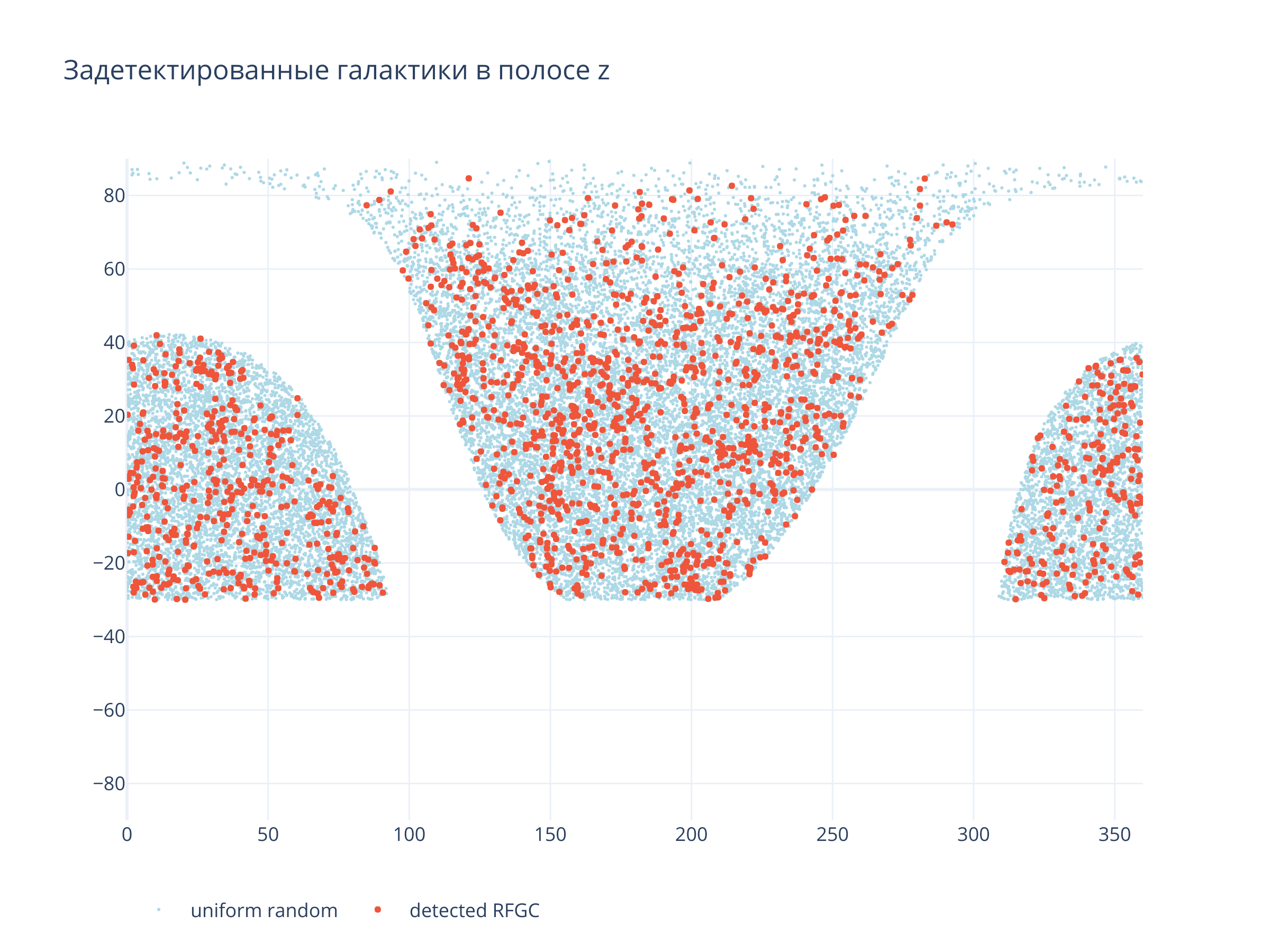
Покрытие неба¶
fig = go.Figure()
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
#width=900, height=700,
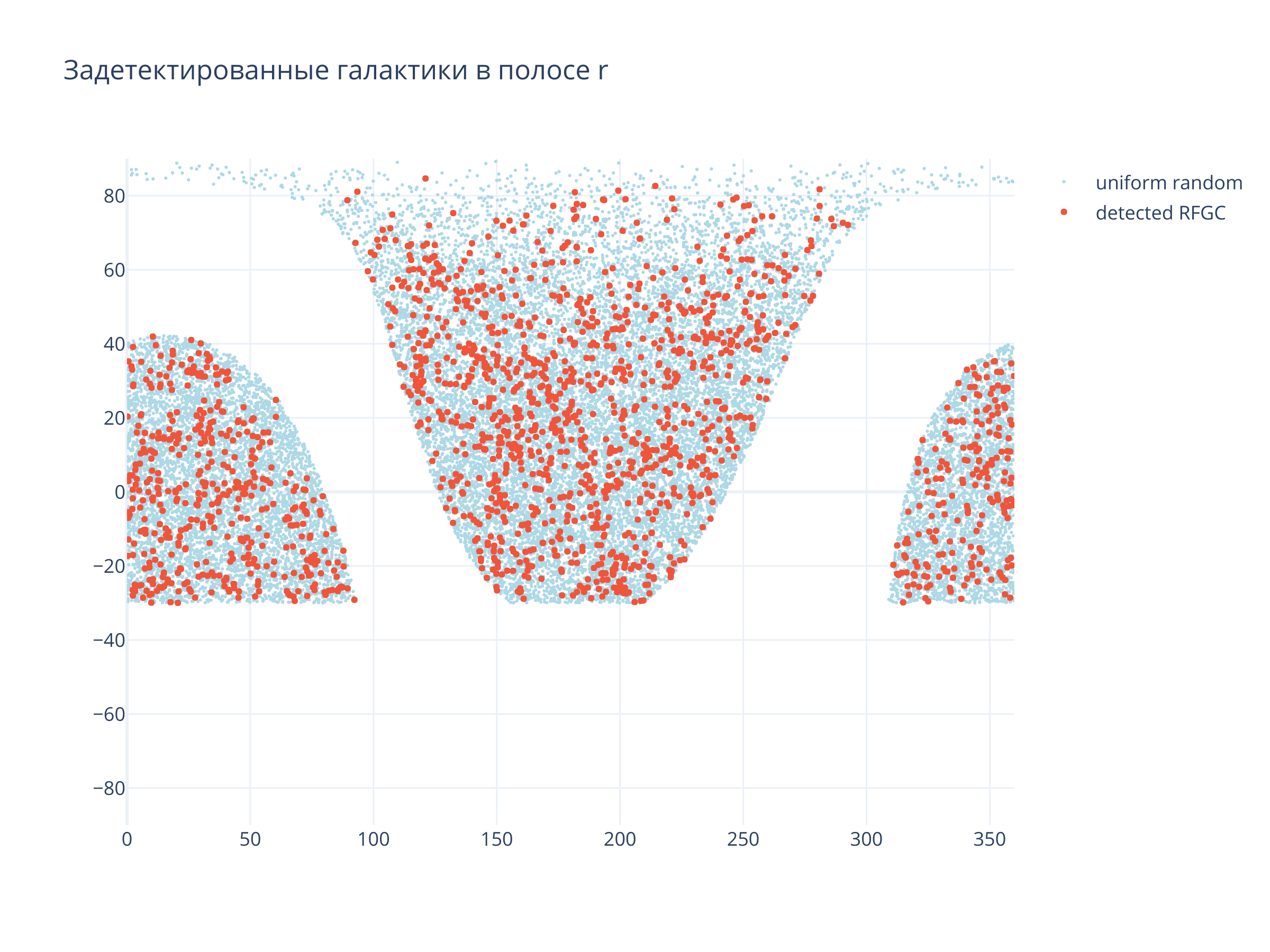
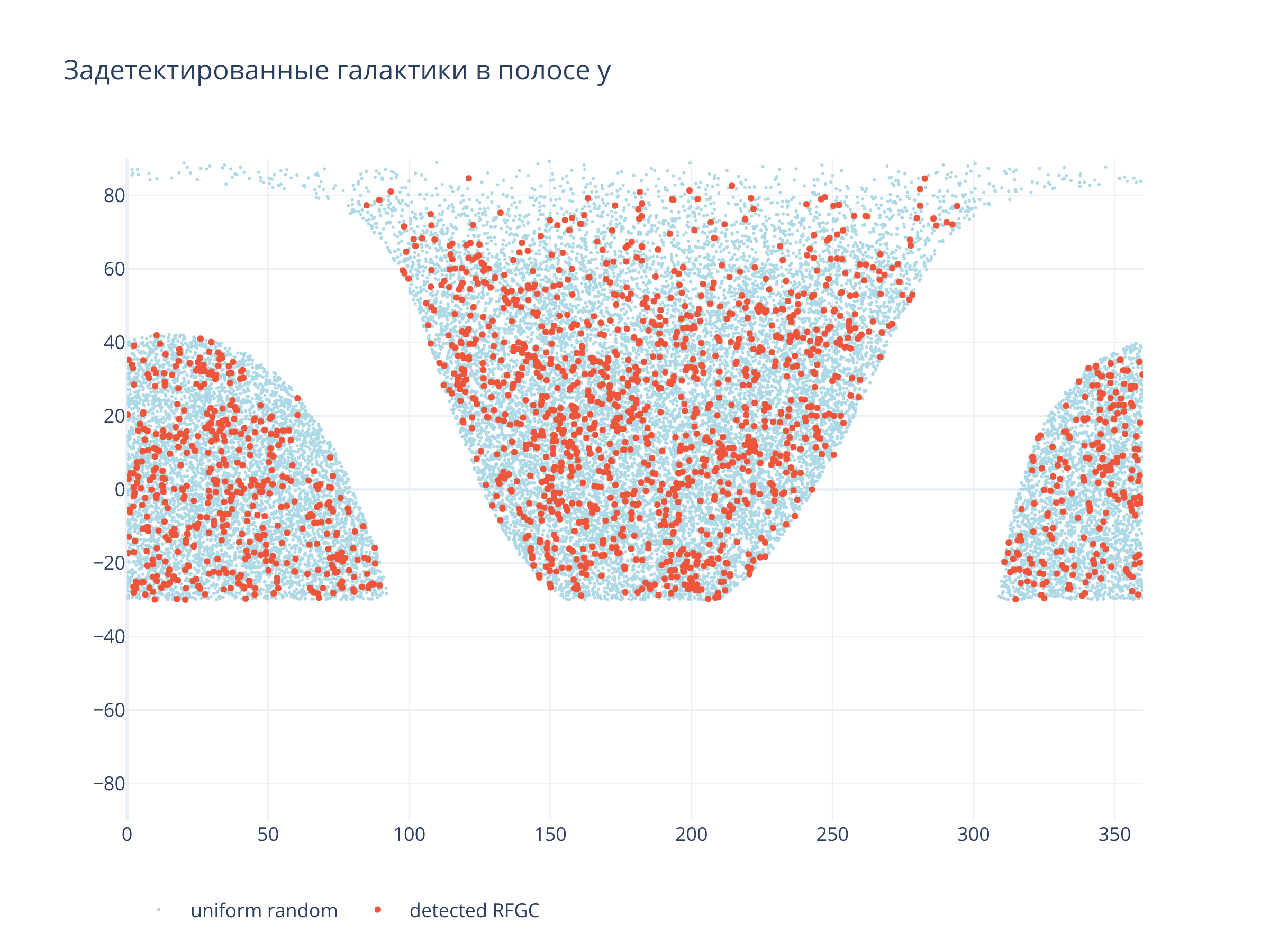
title = f"Задетектированные галактики в полосе {filt}",
legend_orientation = 'h'
)
fig
Светло-синие точки --- равномерно распределённые по области детектирования случайные точки
Выводы¶
nrfgc = len(df.drop_duplicates(subset=['RFGC']))
import os
import getpass
if not os.environ.get('CASJOBS_WSID'):
os.environ['CASJOBS_WSID'] = input('Enter Casjobs WSID (561085547):')
if not os.environ.get('CASJOBS_PW'):
os.environ['CASJOBS_PW'] = getpass.getpass('Enter Casjobs password:')
import mastcasjobs
jobs = mastcasjobs.MastCasJobs(context="PanSTARRS_DR2", request_type='POST')
from code.hmte import expand_templates
count_template = "".join(open(querypath / 'serscount.tsql').readlines())
@memory.cache
def casquickcaching(jobs, query, task_name):
res = jobs.quick(query, task_name=task_name)
return res
@memory.cache
def count_panstarrs(sqlconds, filt):
r = {}
for k,v in sqlconds.items():
whereclause = f"WHERE {v}" if v != "" else ""
query = expand_templates(count_template, filters=filt, serwhere=whereclause)
taskname = f"count stuff where {k}"
try:
r[k] = (casquickcaching(jobs, query, task_name=taskname))[0][0]
except Exception as e:
print(e)
r[k] = -1
#r[k] = (jobs.quick(query, task_name=f"count stuff where {k}"))[0][0]
print(whereclause, r[k])
return r
t = biggestsermag_aligned
def build_summary(t, filt):
summary = {}
for k,v in masks_t[filt].items():
summary[k] = [len(t.loc[v[v].index]), len(t.loc[v[v].index])/nrfgc]
summary = pd.DataFrame(summary).T
summary.columns = ['N', 'N/N0']
summary['N/Nd'] = summary['N'] / (nrfgc * gal_disc_ratio)
summary['PS'] = np.nan
summary.loc[:, 'PS'] = pd.Series(count_panstarrs(sqlconds[filt], filt))
summary.PS = summary.PS.astype(int)
summary.N = summary.N.astype(int)
return summary
filt='g'
summary[filt] = build_summary(biggestsermag_aligned, filt)
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt]
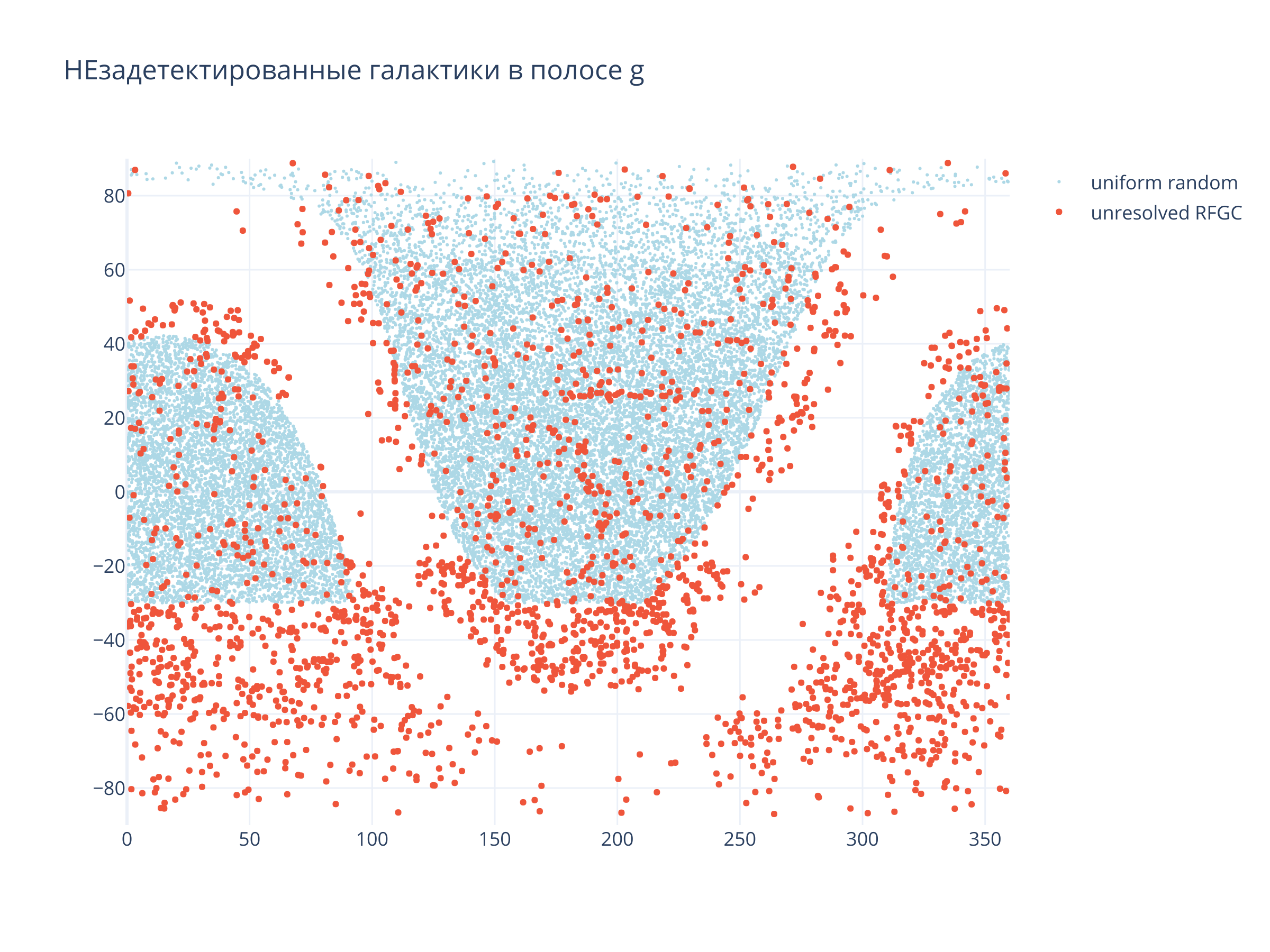
Посмотрим, что не отождествилось вовсе
t1 = df.drop_duplicates(subset=['RFGC'])
t2 = biggestsermag_aligned.drop_duplicates(subset=["RFGC"])
df = pd.concat([t1, t2],sort=True) # concat dataframes
df = df.reset_index(drop=True) # reset the index
df_gpby = df.groupby("RFGC") #group by
idx = [x[0] for x in df_gpby.groups.values() if len(x) == 1] #reindex
unresolved = t1.iloc[idx]
fig = go.Figure()
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = unresolved['RAJ2000'],
y = unresolved['DEJ2000'], name = "unresolved RFGC",
text = list(unresolved['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
title = f"НЕзадетектированные галактики в полосе {filt}"
)
fig
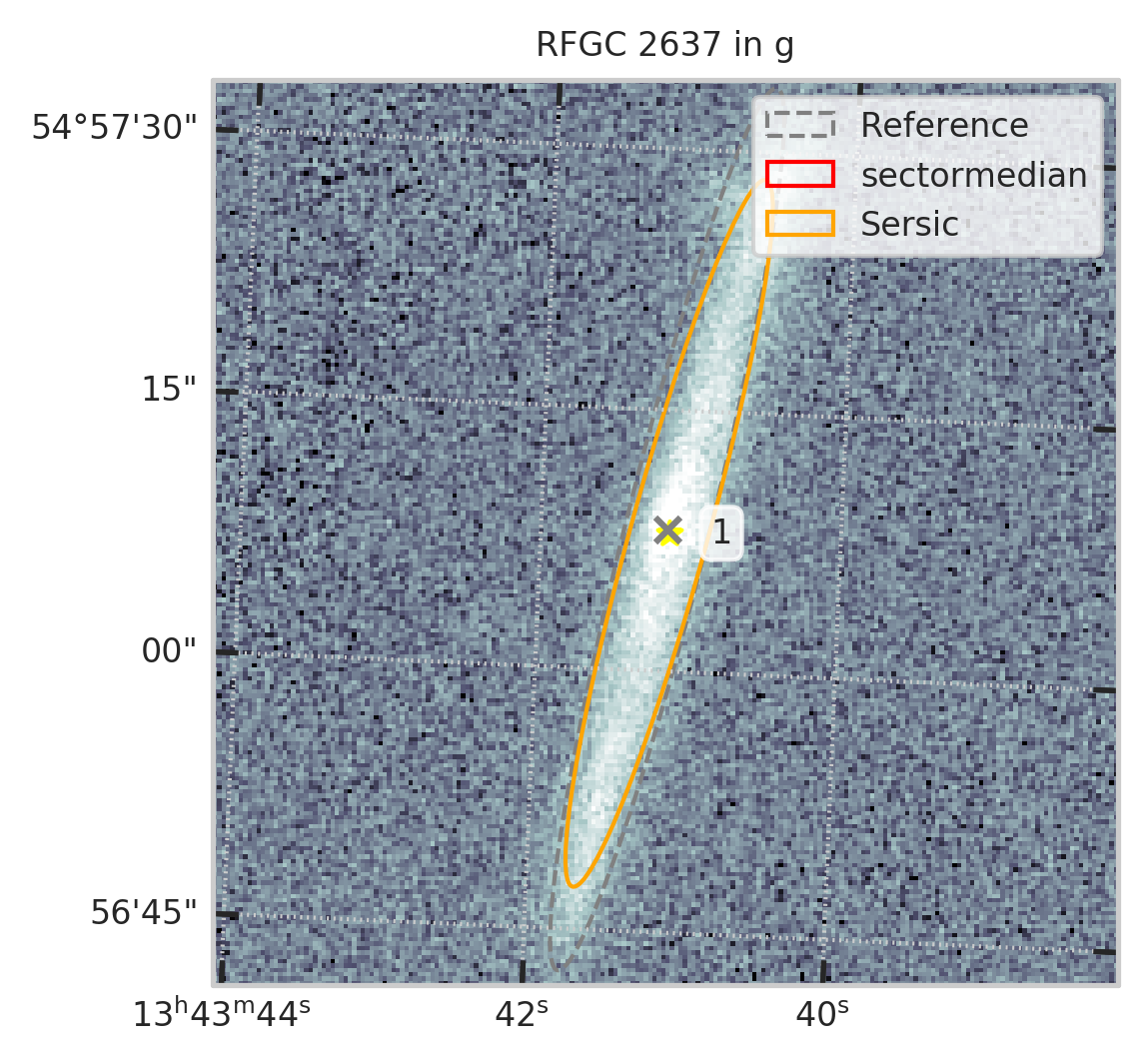
subset = df[df.RFGC == 2637]
plt.figure(figsize=(4,4))
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=1, transform=LogStretch(10) + PercentileInterval(99.5))
plt.show()
with pd.option_context("display.max_rows", 1000): display(subset.T)
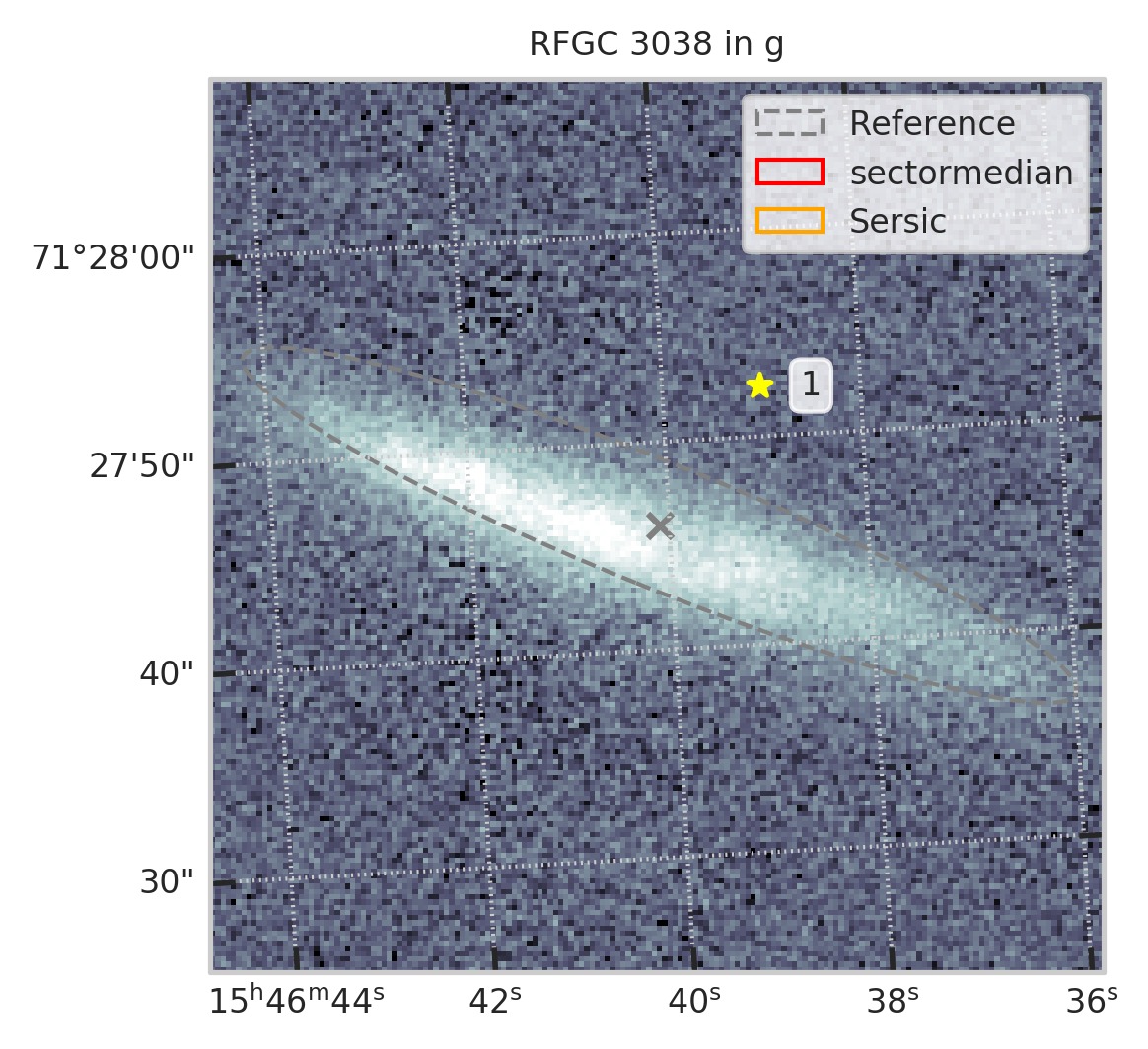
subset = df[df.RFGC == 3038]
plt.figure(figsize=(4,4))
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=1, transform=LogStretch(10) + PercentileInterval(99.5))
plt.show()
with pd.option_context("display.max_rows", 1000): display(subset.T)
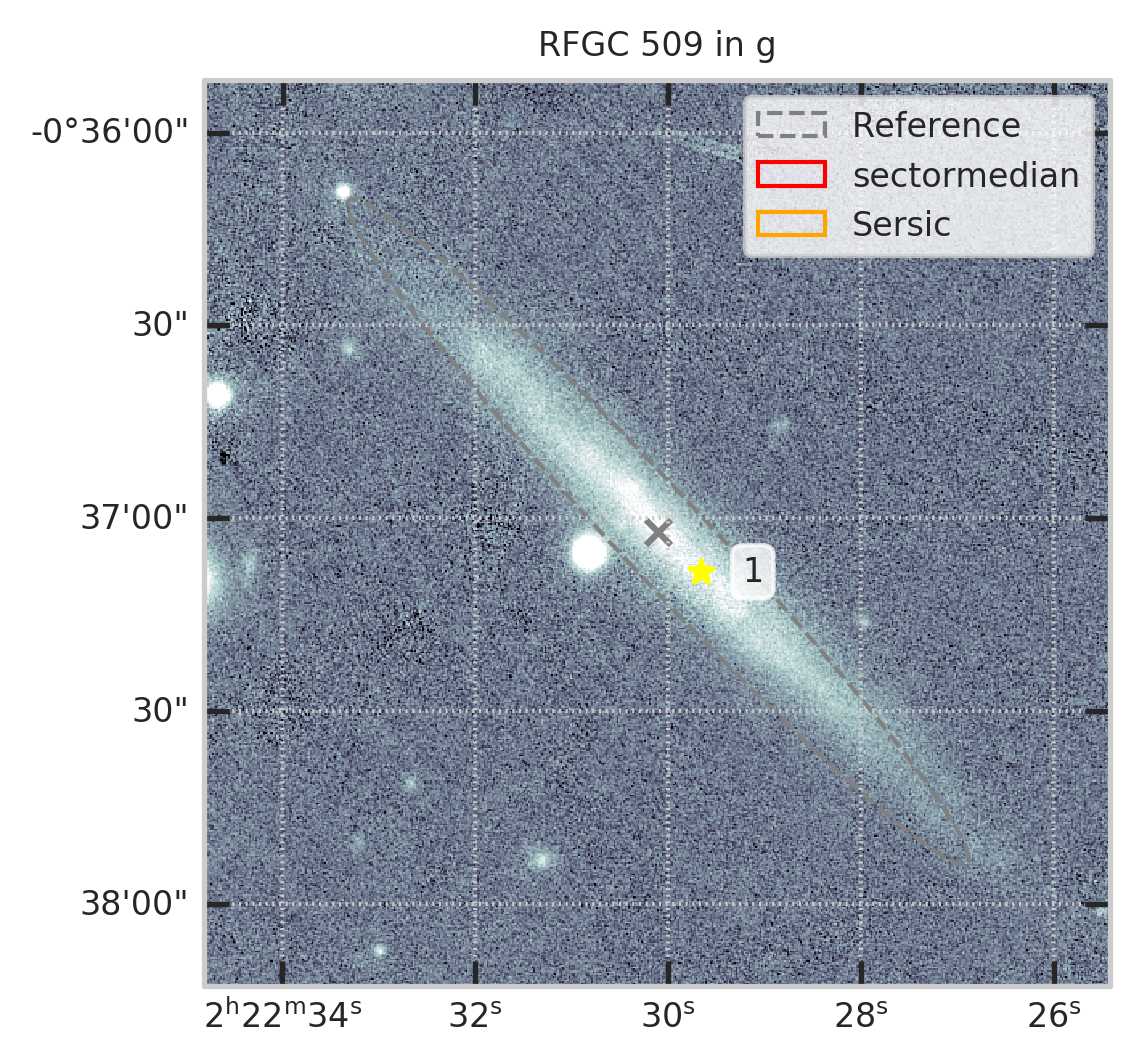
subset = df[df.RFGC == 509]
plt.figure(figsize=(4,4))
show_galaxy_rfgc(subset.iloc[0], 'g', df=subset, image=True, fig=fig,
sersic=True, cleanp=False, kron=False, median=True, petrosian=False, exp=False, voculer=False,
zoom=1, transform=LogStretch(10) + PercentileInterval(99.5))
plt.show()
with pd.option_context("display.max_rows", 1000): display(subset.T)
не нашёл он галактику...................
Подбор критериев отбора (r)¶
filt = 'r'
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex"])
fitted = wshape.dropna(subset=[f"{filt}SerRadius"])
sane = fitted[fitted[f"{filt}SerA"] > 0.051]
aligned = sane[(np.abs(sane[f"{filt}GalPhi"] - sane["PA"]) < 10) & \
(np.abs(sane[f"{filt}SerPhi"] - sane["PA"]) < 10)]
##aligned.index= range(1, len(aligned)+1)
biggestsermag_aligned = aligned.sort_values(["RFGC",f"{filt}SerMag"],ascending=[True, True]).groupby('RFGC').head(1)
t = biggestsermag_aligned
t['BaO'] = 1/t['AbO']
t['AbE'] = t['bE']/t['aE']
t['BaE'] = 1/t['AbE']
from scipy.stats import gaussian_kde
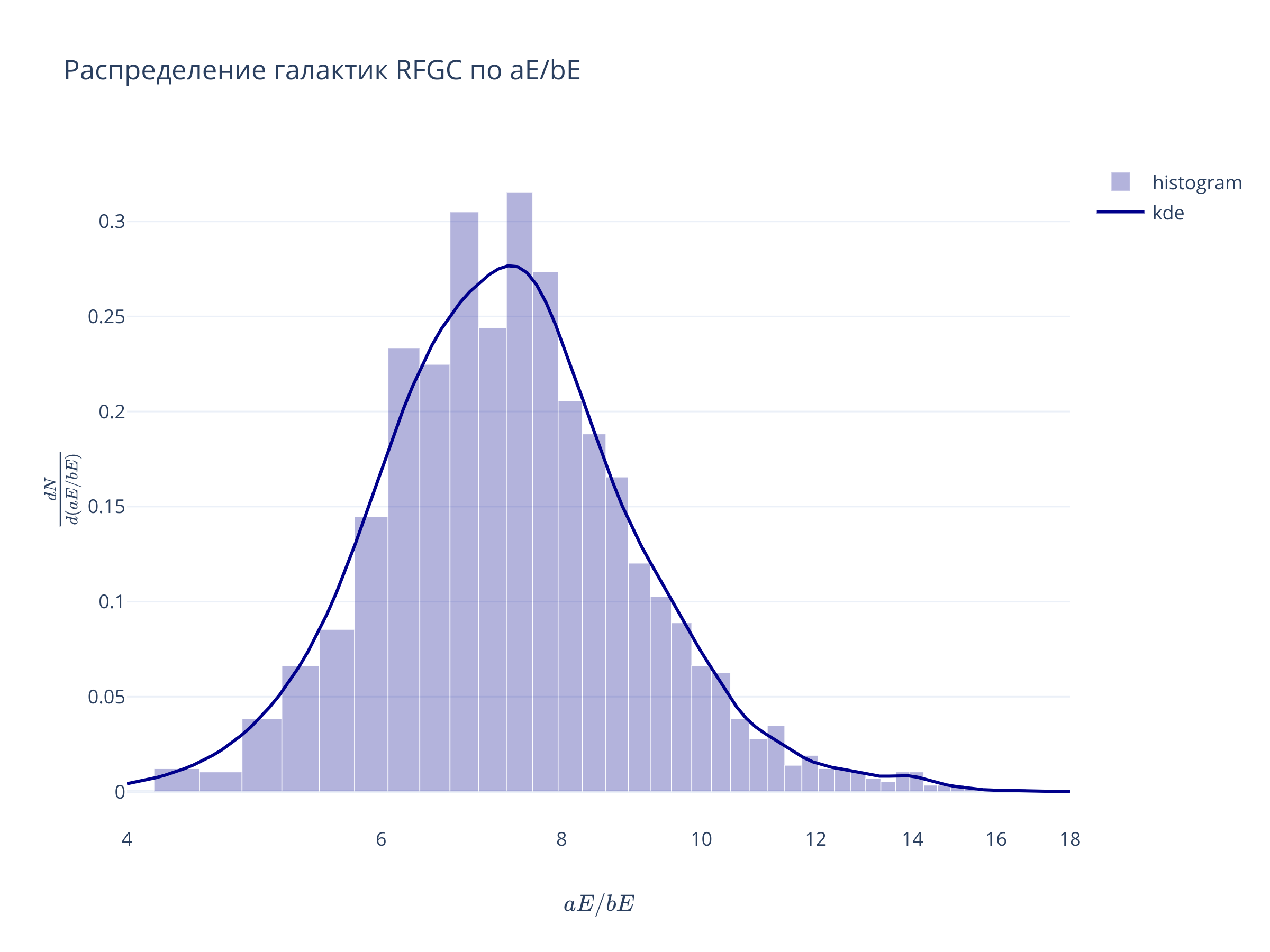
yy, xx_e = np.histogram(t['BaE'], bins='auto', density=True)
xx = (xx_e[1:] + xx_e[:-1])/2
x1, x2 = np.log10(4), np.log10(18)
kernel = gaussian_kde(t['BaE'])
x_grid = np.logspace(x1, x2, 100)
go.Figure(
data = [
go.Bar(x = xx, y = yy, width=xx[1]-xx[0], opacity=0.3, marker = {'color':'darkblue'}, name="histogram"),
go.Scatter(x = x_grid, y = kernel(x_grid), line = {'color':'darkblue'}, name = 'kde')
],
layout = go.Layout(xaxis = {'range':(x1, x2), 'type':'log', 'title':r"$aE/bE$"},
yaxis = {'title':r"$\frac{dN}{d (aE/bE)}$"},
title="Распределение галактик RFGC по aE/bE")
)
def scadensrfgc_ply(t, x, y, xrange, yrange, **kw):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
#print(len(xx))
mode = kw.pop('mode', 'kde')
contours = kw.pop('contours', True)
logx = kw.pop('logx', False)
logy = kw.pop('logy', True)
plots_kw = dict(mode=mode, modepars={'bins':(30,30)}, alpha=0.9,
pointlabels=rfgc_names, contours=contours, **kw)
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=logy, logx=logx,
**plots_kw)
return fig
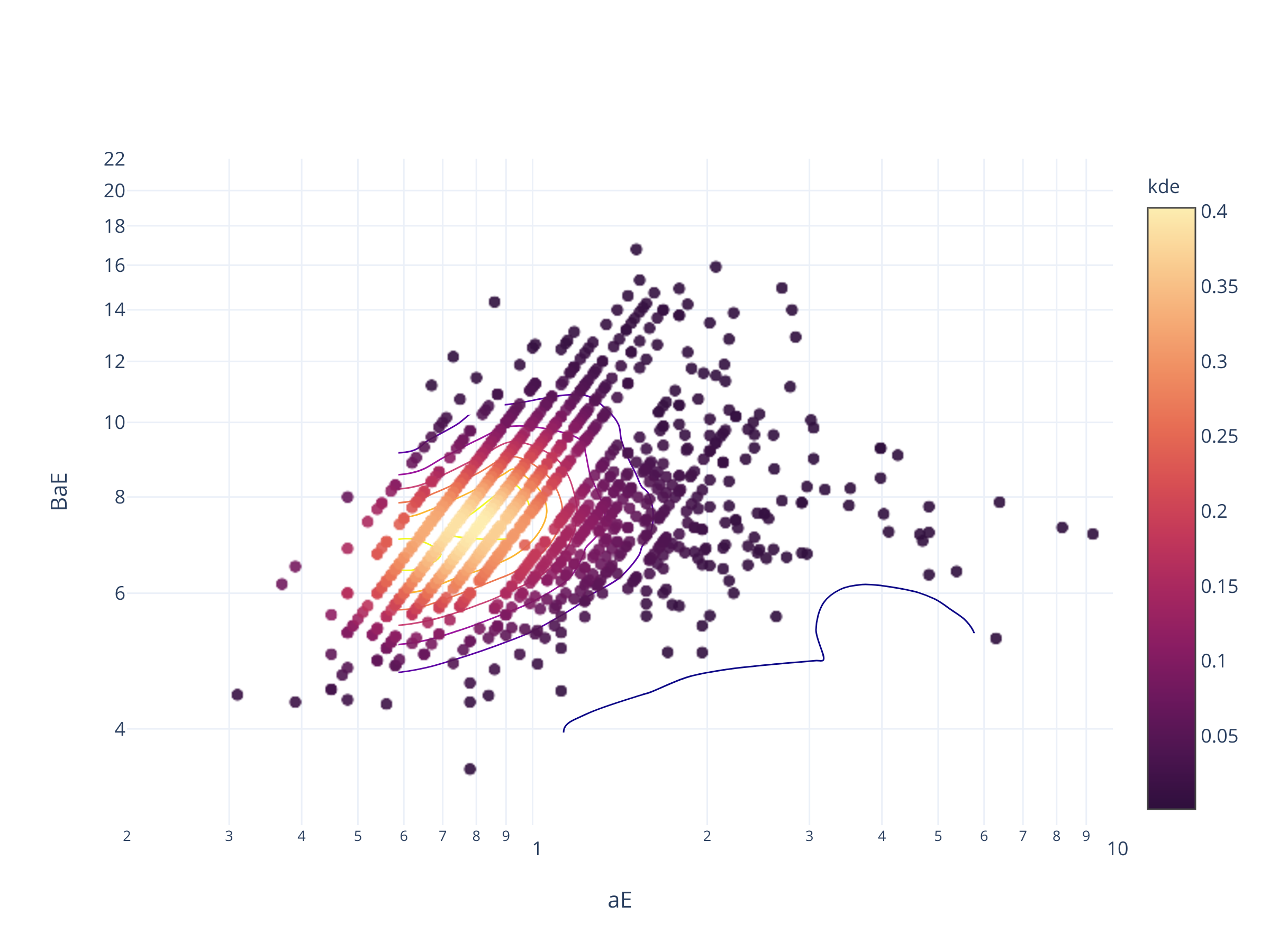
fig = scadensrfgc_ply(
biggestsermag_aligned,
'aE', 'BaE',
(0.2, 10), (3, 22),
logx=True
)
fig.update_layout(autosize=False, width=600, height=500)
fig
Посмотрим как оно может выглядеть для объектов PanSTARRS
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 1/0.043)
)
fig.update_layout(autosize=False, width=600, height=500)
fig
0. Границы по большой полуоси¶
FWHM у PSF в обзоре $1''-2''$ (Глянул у пары галактик, но надо ещё уточнить)
Если у галактики полуось < $7''$, она будет видна с отношением осей $<7$ (а если нет, то это ошибка алгоритма). Получается, что надо отсекать где-то по $5'' - 7''$
t = biggestsermag_aligned
sizemask = {
5:(t[f'{filt}SerA'] > 5),
7:(t[f'{filt}SerA'] > 7),
}
summary[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index = t.index)}
sqlconds[filt] = {'all':f"{filt}SerRadius > 0.051"}
1. По «наблюдаемому» соотношению осей¶
Можно попробовать подобрать границу «круглости» с помощью этой функции
from code.plotutils import scatter_density_plotly, seaborn_to_plotly
def a_b_obs(a, a_b, sigma=1, inverse=False, ):
if inverse:
b = a * (1/a_b)
ratio = np.sqrt(b**2 + sigma**2) / np.sqrt(a**2 + sigma**2)
else:
b = a / a_b
ratio = np.sqrt(a**2 + sigma**2)/np.sqrt(b**2 + sigma**2)
return ratio
def b_a_obs(a, b_a, sigma=1):
return a_b_obs(a, 1/b_a, sigma, inverse=True)
x, y = f'{filt}SerA', f'{filt}SerAb'
y2 = f'{filt}SerBa'
xrange, yrange = (1, 26), (0.043, 1)
t = biggestsermag_aligned
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
sizes = np.linspace(*xrange, 51)
a_b_list = np.array([7, 25])
b_a_list = 1/a_b_list
N = len(a_b_list)
plots_kw = dict(mode="kde", modepars={'bins':(30,30)}, contours=True, alpha=0.9,
pointlabels=rfgc_names)
subplotlayout = dict(rows=1, cols=2)
fig = make_subplots(**subplotlayout)
fig.layout.update(
xaxis = {'domain':[0, 0.4]},
xaxis2 = {'domain':[0.6, 1]},
)
fig.layout.xaxis.type = 'log'
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=True, logx=True,
subplotpos={'row':1, 'col':1}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig = scatter_density_plotly(xx, 1/yy, xrange, (1/yrange[1], 1/yrange[0]), x, y2, logy=True, logx=True,
subplotpos={'row':1, 'col':2}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig.update_traces(
marker = {'colorbar':{'x':0.4}},
selector = {'type':'scattergl'},
row = 1, col = 1)
fig.update_traces(
marker = {'colorbar':{'x':1}},
selector = {'type':'scattergl'},
row = 1, col = 2)
fig.update_layout(showlegend=True, legend_orientation="h")
pallete = sb.cubehelix_palette(4, start=.5, gamma=0.8, rot=-0.7, dark=0.25, light=0.75)
ply_chx = seaborn_to_plotly(pallete)
nncurves = len(fig.data)
for i, a_b in enumerate(a_b_list):
fig.add_trace(
go.Scattergl(
x = sizes,
y = b_a_obs(sizes, 1/a_b),
name="%.2f" % a_b, showlegend=True,
legendgroup="%.2f" % a_b,
hoverinfo='name',
line = {'color':ply_chx[i][1]}
), row=1, col=1)
fig.add_trace(
go.Scattergl(
x = sizes,
y = a_b_obs(sizes, a_b),
name = "%.2f" % a_b,
hoverinfo='name', showlegend=False,
line = {'color':ply_chx[i][1]}
), row=1, col=2)
for i, x in enumerate([5, 7]):
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo = "name", legendgroup=f"a = {x}"),
row=1, col=1)
yrange2 = (1/yrange[1], 1/yrange[0])
fig.add_trace(go.Scattergl(x = [x, x], y = yrange2, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo="name", showlegend=False),
row=1, col=2)
fig.layout.legend.x = 0.3
fig.layout.legend.y = -0.2
fig.layout.legend.tracegroupgap = 30
fig.layout.title = "Распредение объектов по полуоси и отношению осей и наблюдаемое отношение осей"
fw = go.FigureWidget(fig)
@interact(a_b1 = (1, 30, .1), a_b2 = (1, 30, .1), sigma=(1, 10, 0.1))
def update(a_b1 = 7, a_b2 = 18, sigma=3.8):
with fw.batch_update():
for i, (a_b, inverse) in enumerate([(a_b1, True), (a_b1, False), (a_b2, True), (a_b2, False)],
start=nncurves):
fw.data[i].update(
y = a_b_obs(sizes, a_b, sigma, inverse=inverse),
name = "%.2f" % a_b)
fw.show(height=400, width=900)
def sel_obs_ratio(t, r_obs, sigma, amin):
key = f'r_obs={r_obs:d} sigma={sigma:.2f} a>{amin:d}'
masks_t[filt][key] = (
(t[f'{filt}SerAb'] < a_b_obs(t[f'{filt}SerA'], r_obs, sigma=sigma, inverse=True)) &
sizemask[amin]
)
sqlconds[filt][key] = f"{filt}SerAb < SQRT(SQUARE({filt}SerRadius/{r_obs:.2f}) + {sigma**2:.2f}) / SQRT(SQUARE({filt}SerRadius) + {sigma**2:.2f}) AND {filt}SerRadius > {amin:.2f}"
sel_obs_ratio(biggestsermag_aligned, 7, 3.8, 5)
sel_obs_ratio(biggestsermag_aligned, 7, 3.8, 7)
2. По дисперсии линейной регрессии¶
import statsmodels.api as sm
from astropy.modeling import models, fitting
def compute_linregr_pars(t, x, y, xrange, yrange, logx=False, logy=True):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
xx_reg = (t.loc[mask, x])
yy_reg = (t.loc[mask, y])
xx_reg = np.log10(xx_reg) if logx else xx_reg
yy_reg = np.log10(yy_reg) if logy else yy_reg
variables = pd.DataFrame({
'const':np.ones_like(xx_reg),
'x':xx_reg,
#'x2':xx**2,
})
linmodel = sm.OLS(yy_reg, variables)
a,b,c,d,e = [0]*5
mod = Namespace()
linresults = linmodel.fit()
mod.eps = linresults.resid
mod.b,mod.a = linresults.params
hist, xe = np.histogram(mod.eps)
xe = (xe[1:]+xe[:-1])/2
ini = models.Gaussian1D()
fitter = fitting.LevMarLSQFitter()
fit = fitter(ini, xe, hist)
mod.sigma_total = fit.stddev
return mod
xrange, yrange = (1, 26), (1, 23.26)
m = compute_linregr_pars(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
x_grid = np.linspace(*xrange, 100)
fitted = 10**(m.a*x_grid + m.b)
upperbnd = 10**(m.a*x_grid + m.b + 2*m.sigma_total)
lowerbnd = 10**(m.a*x_grid + m.b - 2*m.sigma_total)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
fig.add_trace(go.Scattergl(x = x_grid, y = fitted, name='fitted'))
fig.add_trace(go.Scattergl(x = np.hstack([x_grid, [x_grid[-1]+1,x_grid[-1]+1], np.flip(x_grid)]),
y = np.hstack([upperbnd, [upperbnd[-1], lowerbnd[-1]], np.flip(lowerbnd)]), line = {'color':'lightgray', 'width':3},
name='2σ range'))
fig.layout.legend.x = 1.3
fig.update_layout(
title = "Линейная регрессия и диапазон в 2σ",
clickmode = "event+select",
autosize=False,width=700, height=500)
fig.show()
def sel_lbnd_linreg(t, amin):
key = f'2 sigma a>{amin:d}'
masks_t[filt][key] = (
(np.log10(t[f'{filt}SerBa']) > m.a*t[f'{filt}SerA']+m.b - 2*m.sigma_total) &
(t[f'{filt}SerA'] > amin)
)
sqlconds[filt][key] = f"-LOG10({filt}SerAb) > {m.a:.5f}*gSerRadius + {m.b:.5f} - {2*m.sigma_total:.5f} AND {filt}SerRadius > {amin:.2f}"
#t = biggestsermag_aligned
sel_lbnd_linreg(biggestsermag_aligned, 5)
sel_lbnd_linreg(biggestsermag_aligned, 7)
3. Простые границы¶
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 23.26)
)
x_grid = np.linspace(*xrange, 100)
fig.layout.legend.x = 1.3
fig.layout.legend.tracegroupgap = 20
fig.update_layout(
title = "Простые границы по отношению",
clickmode = "event+select",
autosize=False,width=800, height=600)
for x in [5, 7]:
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", name=f"a = {x}", legendgroup=f"a = {x}"))
for y in [3, 4]:
fig.add_trace(go.Scattergl(x = xrange, y = [y,y],mode = "lines",name=f"a/b = {y}", legendgroup = f"a/b = {y}"))
fig
def sel_tresh(t, ratio, amin):
key = f'a/b>{ratio:d} a>{amin:d}'
masks_t[filt][key] = ((t[f'{filt}SerA'] > amin) & (t[f'{filt}SerBa'] > ratio))
sqlconds[filt][key] = f"{filt}SerRadius > {amin:.2f} AND {filt}SerAb < {1/ratio:.5f}"
sel_tresh(biggestsermag_aligned, 3, 5)
sel_tresh(biggestsermag_aligned, 3, 7)
sel_tresh(biggestsermag_aligned, 4, 5)
sel_tresh(biggestsermag_aligned, 4, 7)
Покрытие неба¶
fig = go.Figure()
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
title = f"Задетектированные галактики в полосе {filt}"
)
fig
Светло-синие точки --- равномерно распределённые по области детектирования случайные точки
Выводы¶
summary[filt] = build_summary(biggestsermag_aligned, filt)
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt]
Подбор критериев отбора (i)¶
filt='i'
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex"])
fitted = wshape.dropna(subset=[f"{filt}SerRadius"])
sane = fitted[fitted[f"{filt}SerA"] > 0.051]
aligned = sane[(np.abs(sane[f"{filt}GalPhi"] - sane["PA"]) < 10) & \
(np.abs(sane[f"{filt}SerPhi"] - sane["PA"]) < 10)]
#aligned.index= range(1, len(aligned)+1)
biggestsermag_aligned = aligned.sort_values(["RFGC",f"{filt}SerMag"],ascending=[True, True]).groupby('RFGC').head(1)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 1/0.043)
)
fig.update_layout(autosize=False, width=600, height=500)
fig
0. Границы по большой полуоси¶
FWHM у PSF в обзоре $1''-2''$ (Глянул у пары галактик, но надо ещё уточнить)
Если у галактики полуось < $7''$, она будет видна с отношением осей $<7$ (а если нет, то это ошибка алгоритма). Получается, что надо отсекать где-то по $5'' - 7''$
t = biggestsermag_aligned
sizemask = {
5:(t[f'{filt}SerA'] > 5),
7:(t[f'{filt}SerA'] > 7),
}
summary[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index=t.index)}
sqlconds[filt] = {'all':f"{filt}SerRadius > 0.051"}
1. По «наблюдаемому» соотношению осей¶
Можно попробовать подобрать границу «круглости» с помощью этой функции
from code.plotutils import scatter_density_plotly, seaborn_to_plotly
x, y = f'{filt}SerA', f'{filt}SerAb'
y2 = f'{filt}SerBa'
xrange, yrange = (1, 30), (0.043, 1)
t = biggestsermag_aligned
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
sizes = np.linspace(*xrange, 51)
a_b_list = np.array([7, 25])
b_a_list = 1/a_b_list
N = len(a_b_list)
plots_kw = dict(mode="kde", modepars={'bins':(30,30)}, contours=True, alpha=0.9,
pointlabels=rfgc_names)
subplotlayout = dict(rows=1, cols=2)
fig = make_subplots(**subplotlayout)
fig.layout.update(
xaxis = {'domain':[0, 0.4]},
xaxis2 = {'domain':[0.6, 1]},
)
fig.layout.xaxis.type = 'log'
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=True, logx=True,
subplotpos={'row':1, 'col':1}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig = scatter_density_plotly(xx, 1/yy, xrange, (1/yrange[1], 1/yrange[0]), x, y2, logy=True, logx=True,
subplotpos={'row':1, 'col':2}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig.update_traces(
marker = {'colorbar':{'x':0.4}},
selector = {'type':'scattergl'},
row = 1, col = 1)
fig.update_traces(
marker = {'colorbar':{'x':1}},
selector = {'type':'scattergl'},
row = 1, col = 2)
fig.update_layout(showlegend=True, legend_orientation="h")
pallete = sb.cubehelix_palette(4, start=.5, gamma=0.8, rot=-0.7, dark=0.25, light=0.75)
ply_chx = seaborn_to_plotly(pallete)
nncurves = len(fig.data)
for i, a_b in enumerate(a_b_list):
fig.add_trace(
go.Scattergl(
x = sizes,
y = b_a_obs(sizes, 1/a_b),
name="%.2f" % a_b, showlegend=True,
legendgroup="%.2f" % a_b,
hoverinfo='name',
line = {'color':ply_chx[i][1]}
), row=1, col=1)
fig.add_trace(
go.Scattergl(
x = sizes,
y = a_b_obs(sizes, a_b),
name = "%.2f" % a_b,
hoverinfo='name', showlegend=False,
line = {'color':ply_chx[i][1]}
), row=1, col=2)
for i, x in enumerate([5, 7]):
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo = "name", legendgroup=f"a = {x}"),
row=1, col=1)
yrange2 = (1/yrange[1], 1/yrange[0])
fig.add_trace(go.Scattergl(x = [x, x], y = yrange2, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo="name", showlegend=False),
row=1, col=2)
fig.layout.legend.x = 0.3
fig.layout.legend.y = -0.2
fig.layout.legend.tracegroupgap = 30
fig.layout.title = "Распредение объектов по полуоси и отношению осей и наблюдаемое отношение осей"
fw = go.FigureWidget(fig)
@interact(a_b1 = (1, 30, .1), a_b2 = (1, 30, .1), sigma=(1, 10, 0.1))
def update(a_b1 = 7, a_b2 = 18, sigma=3.5):
with fw.batch_update():
for i, (a_b, inverse) in enumerate([(a_b1, True), (a_b1, False), (a_b2, True), (a_b2, False)],
start=nncurves):
fw.data[i].update(
y = a_b_obs(sizes, a_b, sigma, inverse=inverse),
name = "%.2f" % a_b)
fw.show(height=400, width=900)
def sel_obs_ratio(t, r_obs, sigma, amin):
key = f'r_obs={r_obs:d} sigma={sigma:.2f} a>{amin:d}'
masks_t[filt][key] = (
(t[f'{filt}SerAb'] < a_b_obs(t[f'{filt}SerA'], r_obs, sigma=sigma, inverse=True)) &
sizemask[amin]
)
sqlconds[filt][key] = f"{filt}SerAb < SQRT(SQUARE({filt}SerRadius/{r_obs:.2f}) + {sigma**2:.2f}) / SQRT(SQUARE({filt}SerRadius) + {sigma**2:.2f}) AND {filt}SerRadius > {amin:.2f}"
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 5)
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 7)
2. По дисперсии линейной регрессии¶
import statsmodels.api as sm
from astropy.modeling import models, fitting
def compute_linregr_pars(t, x, y, xrange, yrange, logx=False, logy=True):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
xx_reg = (t.loc[mask, x])
yy_reg = (t.loc[mask, y])
xx_reg = np.log10(xx_reg) if logx else xx_reg
yy_reg = np.log10(yy_reg) if logy else yy_reg
variables = pd.DataFrame({
'const':np.ones_like(xx_reg),
'x':xx_reg,
#'x2':xx**2,
})
linmodel = sm.OLS(yy_reg, variables)
a,b,c,d,e = [0]*5
mod = Namespace()
linresults = linmodel.fit()
mod.eps = linresults.resid
mod.b,mod.a = linresults.params
hist, xe = np.histogram(mod.eps)
xe = (xe[1:]+xe[:-1])/2
ini = models.Gaussian1D()
fitter = fitting.LevMarLSQFitter()
fit = fitter(ini, xe, hist)
mod.sigma_total = fit.stddev
return mod
xrange, yrange = (1, 26), (1, 23.26)
m = compute_linregr_pars(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
x_grid = np.linspace(*xrange, 100)
fitted = 10**(m.a*x_grid + m.b)
upperbnd = 10**(m.a*x_grid + m.b + 2*m.sigma_total)
lowerbnd = 10**(m.a*x_grid + m.b - 2*m.sigma_total)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
fig.add_trace(go.Scattergl(x = x_grid, y = fitted, name='fitted'))
fig.add_trace(go.Scattergl(x = np.hstack([x_grid, [x_grid[-1]+1,x_grid[-1]+1], np.flip(x_grid)]),
y = np.hstack([upperbnd, [upperbnd[-1], lowerbnd[-1]], np.flip(lowerbnd)]), line = {'color':'lightgray', 'width':3},
name='2σ range'))
fig.layout.legend.x = 1.3
fig.update_layout(
title = "Линейная регрессия и диапазон в 2σ",
clickmode = "event+select",
autosize=False,width=700, height=500)
fig.show()
def sel_lbnd_linreg(t, amin):
key = f'2 sigma a>{amin:d}'
masks_t[filt][key] = (
(np.log10(t[f'{filt}SerBa']) > m.a*t[f'{filt}SerA']+m.b - 2*m.sigma_total) &
(t[f'{filt}SerA'] > amin)
)
sqlconds[filt][key] = f"-LOG10({filt}SerAb) > {m.a:.5f}*gSerRadius + {m.b:.5f} - {2*m.sigma_total:.5f} AND {filt}SerRadius > {amin:.2f}"
#t = biggestsermag_aligned
sel_lbnd_linreg(biggestsermag_aligned, 5)
sel_lbnd_linreg(biggestsermag_aligned, 7)
3. Простые границы¶
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 23.26)
)
x_grid = np.linspace(*xrange, 100)
fig.layout.legend.x = 1.3
fig.layout.legend.tracegroupgap = 20
fig.update_layout(
title = "Простые границы по отношению",
clickmode = "event+select",
autosize=False,width=800, height=600)
for x in [5, 7]:
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", name=f"a = {x}", legendgroup=f"a = {x}"))
for y in [3, 4]:
fig.add_trace(go.Scattergl(x = xrange, y = [y,y],mode = "lines",name=f"a/b = {y}", legendgroup = f"a/b = {y}"))
fig
def sel_tresh(t, ratio, amin):
key = f'a/b>{ratio:d} a>{amin:d}'
masks_t[filt][key] = (t[f'{filt}SerA'] > amin) & (t[f'{filt}SerBa'] > ratio)
sqlconds[filt][key] = f"{filt}SerRadius > {amin:.2f} AND {filt}SerAb < {1/ratio:.5f}"
sel_tresh(biggestsermag_aligned, 3, 5)
sel_tresh(biggestsermag_aligned, 3, 7)
sel_tresh(biggestsermag_aligned, 4, 5)
sel_tresh(biggestsermag_aligned, 4, 7)
Покрытие неба¶
fig = go.Figure()
t = biggestsermag_aligned
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
title = f"Задетектированные галактики в полосе {filt}",
legend_orientation = 'h'
)
fig
Светло-синие точки --- равномерно распределённые по области детектирования случайные точки
Выводы¶
filt = 'i'
summary[filt] = build_summary(biggestsermag_aligned, filt)
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt]
Подбор критериев отбора (z)¶
filt='z'
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex"])
fitted = wshape.dropna(subset=[f"{filt}SerRadius"])
sane = fitted[fitted[f"{filt}SerA"] > 0.051]
aligned = sane[(np.abs(sane[f"{filt}GalPhi"] - sane["PA"]) < 10) & \
(np.abs(sane[f"{filt}SerPhi"] - sane["PA"]) < 10)]
#aligned.index= range(1, len(aligned)+1)
biggestsermag_aligned = aligned.sort_values(["RFGC",f"{filt}SerMag"],ascending=[True, True]).groupby('RFGC').head(1)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 1/0.043)
)
fig.update_layout(autosize=False, width=600, height=500)
fig
0. Границы по большой полуоси¶
FWHM у PSF в обзоре $1''-2''$ (Глянул у пары галактик, но надо ещё уточнить)
Если у галактики полуось < $7''$, она будет видна с отношением осей $<7$ (а если нет, то это ошибка алгоритма). Получается, что надо отсекать где-то по $5'' - 7''$
t = biggestsermag_aligned
sizemask = {
5:(t[f'{filt}SerA'] > 5),
7:(t[f'{filt}SerA'] > 7),
}
summary[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index=t.index)}
sqlconds[filt] = {'all':f"{filt}SerRadius > 0.051"}
1. По «наблюдаемому» соотношению осей¶
Можно попробовать подобрать границу «круглости» с помощью этой функции
from code.plotutils import scatter_density_plotly, seaborn_to_plotly
x, y = f'{filt}SerA', f'{filt}SerAb'
y2 = f'{filt}SerBa'
xrange, yrange = (1, 26), (0.043, 1)
t = biggestsermag_aligned
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
sizes = np.linspace(*xrange, 51)
a_b_list = np.array([7, 25])
b_a_list = 1/a_b_list
N = len(a_b_list)
plots_kw = dict(mode="kde", modepars={'bins':(30,30)}, contours=True, alpha=0.9,
pointlabels=rfgc_names)
subplotlayout = dict(rows=1, cols=2)
fig = make_subplots(**subplotlayout)
fig.layout.update(
xaxis = {'domain':[0, 0.4]},
xaxis2 = {'domain':[0.6, 1]},
)
fig.layout.xaxis.type = 'log'
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=True, logx=True,
subplotpos={'row':1, 'col':1}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig = scatter_density_plotly(xx, 1/yy, xrange, (1/yrange[1], 1/yrange[0]), x, y2, logy=True, logx=True,
subplotpos={'row':1, 'col':2}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig.update_traces(
marker = {'colorbar':{'x':0.4}},
selector = {'type':'scattergl'},
row = 1, col = 1)
fig.update_traces(
marker = {'colorbar':{'x':1}},
selector = {'type':'scattergl'},
row = 1, col = 2)
fig.update_layout(showlegend=True, legend_orientation="h")
pallete = sb.cubehelix_palette(4, start=.5, gamma=0.8, rot=-0.7, dark=0.25, light=0.75)
ply_chx = seaborn_to_plotly(pallete)
nncurves = len(fig.data)
for i, a_b in enumerate(a_b_list):
fig.add_trace(
go.Scatter(
x = sizes,
y = b_a_obs(sizes, 1/a_b),
name="%.2f" % a_b, showlegend=True,
legendgroup="%.2f" % a_b,
hoverinfo='name',
line = {'color':ply_chx[i][1]}
), row=1, col=1)
fig.add_trace(
go.Scatter(
x = sizes,
y = a_b_obs(sizes, a_b),
name = "%.2f" % a_b,
hoverinfo='name', showlegend=False,
line = {'color':ply_chx[i][1]}
), row=1, col=2)
for i, x in enumerate([5, 7]):
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo = "name", legendgroup=f"a = {x}"),
row=1, col=1)
yrange2 = (1/yrange[1], 1/yrange[0])
fig.add_trace(go.Scattergl(x = [x, x], y = yrange2, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo="name", showlegend=False),
row=1, col=2)
fig.layout.legend.x = 0.3
fig.layout.legend.y = -0.2
fig.layout.legend.tracegroupgap = 30
fig.layout.title = "Распредение объектов по полуоси и отношению осей и наблюдаемое отношение осей"
fw = go.FigureWidget(fig)
@interact(a_b1 = (1, 30, .1), a_b2 = (1, 30, .1), sigma=(1, 10, 0.1))
def update(a_b1 = 7, a_b2 = 18, sigma=3.5):
with fw.batch_update():
for i, (a_b, inverse) in enumerate([(a_b1, True), (a_b1, False), (a_b2, True), (a_b2, False)],
start=nncurves):
fw.data[i].update(
y = a_b_obs(sizes, a_b, sigma, inverse=inverse),
name = "%.2f" % a_b)
fw.show(height=400, width=900)
def sel_obs_ratio(t, r_obs, sigma, amin):
key = f'r_obs={r_obs:d} sigma={sigma:.2f} a>{amin:d}'
masks_t[filt][key] = (
(t[f'{filt}SerAb'] < a_b_obs(t[f'{filt}SerA'], r_obs, sigma=sigma, inverse=True)) &
sizemask[amin]
)
sqlconds[filt][key] = f"{filt}SerAb < SQRT(SQUARE({filt}SerRadius/{r_obs:.2f}) + {sigma**2:.2f}) / SQRT(SQUARE({filt}SerRadius) + {sigma**2:.2f}) AND {filt}SerRadius > {amin:.2f}"
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 5)
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 7)
2. По дисперсии линейной регрессии¶
import statsmodels.api as sm
from astropy.modeling import models, fitting
def compute_linregr_pars(t, x, y, xrange, yrange, logx=False, logy=True):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
xx_reg = (t.loc[mask, x])
yy_reg = (t.loc[mask, y])
xx_reg = np.log10(xx_reg) if logx else xx_reg
yy_reg = np.log10(yy_reg) if logy else yy_reg
variables = pd.DataFrame({
'const':np.ones_like(xx_reg),
'x':xx_reg,
#'x2':xx**2,
})
linmodel = sm.OLS(yy_reg, variables)
a,b,c,d,e = [0]*5
mod = Namespace()
linresults = linmodel.fit()
mod.eps = linresults.resid
mod.b,mod.a = linresults.params
hist, xe = np.histogram(mod.eps)
xe = (xe[1:]+xe[:-1])/2
ini = models.Gaussian1D()
fitter = fitting.LevMarLSQFitter()
fit = fitter(ini, xe, hist)
mod.sigma_total = fit.stddev
return mod
xrange, yrange = (1, 26), (1, 23.26)
m = compute_linregr_pars(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
x_grid = np.linspace(*xrange, 100)
fitted = 10**(m.a*x_grid + m.b)
upperbnd = 10**(m.a*x_grid + m.b + 2*m.sigma_total)
lowerbnd = 10**(m.a*x_grid + m.b - 2*m.sigma_total)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
fig.add_trace(go.Scattergl(x = x_grid, y = fitted, name='fitted'))
fig.add_trace(go.Scattergl(x = np.hstack([x_grid, [x_grid[-1]+1,x_grid[-1]+1], np.flip(x_grid)]),
y = np.hstack([upperbnd, [upperbnd[-1], lowerbnd[-1]], np.flip(lowerbnd)]), line = {'color':'lightgray', 'width':3},
name='2σ range'))
fig.layout.legend.x = 1.3
fig.update_layout(
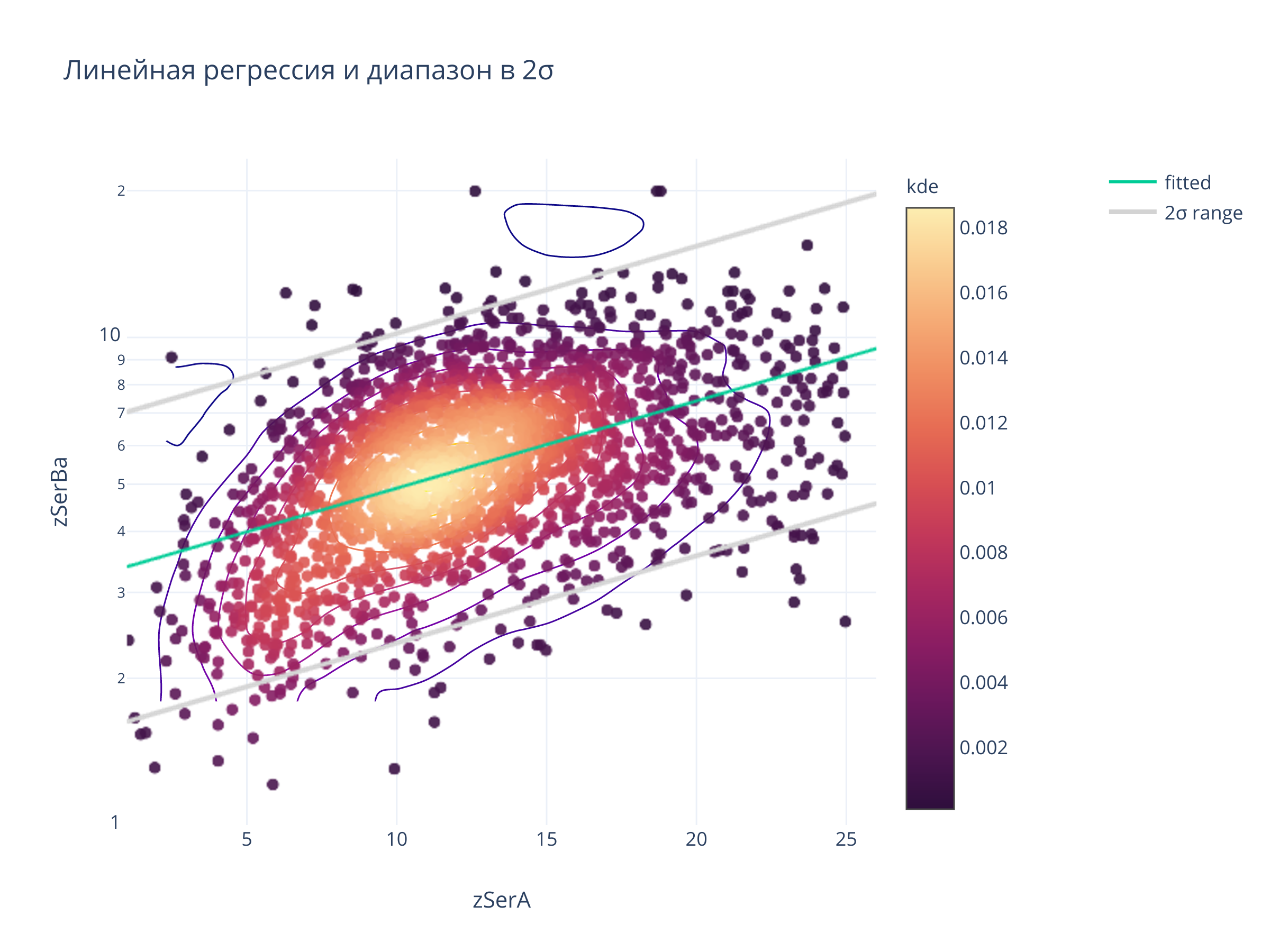
title = "Линейная регрессия и диапазон в 2σ",
clickmode = "event+select",
autosize=False,width=700, height=500)
fig.show()
def sel_lbnd_linreg(t, amin):
key = f'2 sigma a>{amin:d}'
masks_t[filt][key] = (
(np.log10(t[f'{filt}SerBa']) > m.a*t[f'{filt}SerA']+m.b - 2*m.sigma_total) &
(t[f'{filt}SerA'] > amin)
)
sqlconds[filt][key] = f"-LOG10({filt}SerAb) > {m.a:.5f}*gSerRadius + {m.b:.5f} - {2*m.sigma_total:.5f} AND {filt}SerRadius > {amin:.2f}"
#t = biggestsermag_aligned
sel_lbnd_linreg(biggestsermag_aligned, 5)
sel_lbnd_linreg(biggestsermag_aligned, 7)
3. Простые границы¶
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 23.26)
)
x_grid = np.linspace(*xrange, 100)
fig.layout.legend.x = 1.3
fig.layout.legend.tracegroupgap = 20
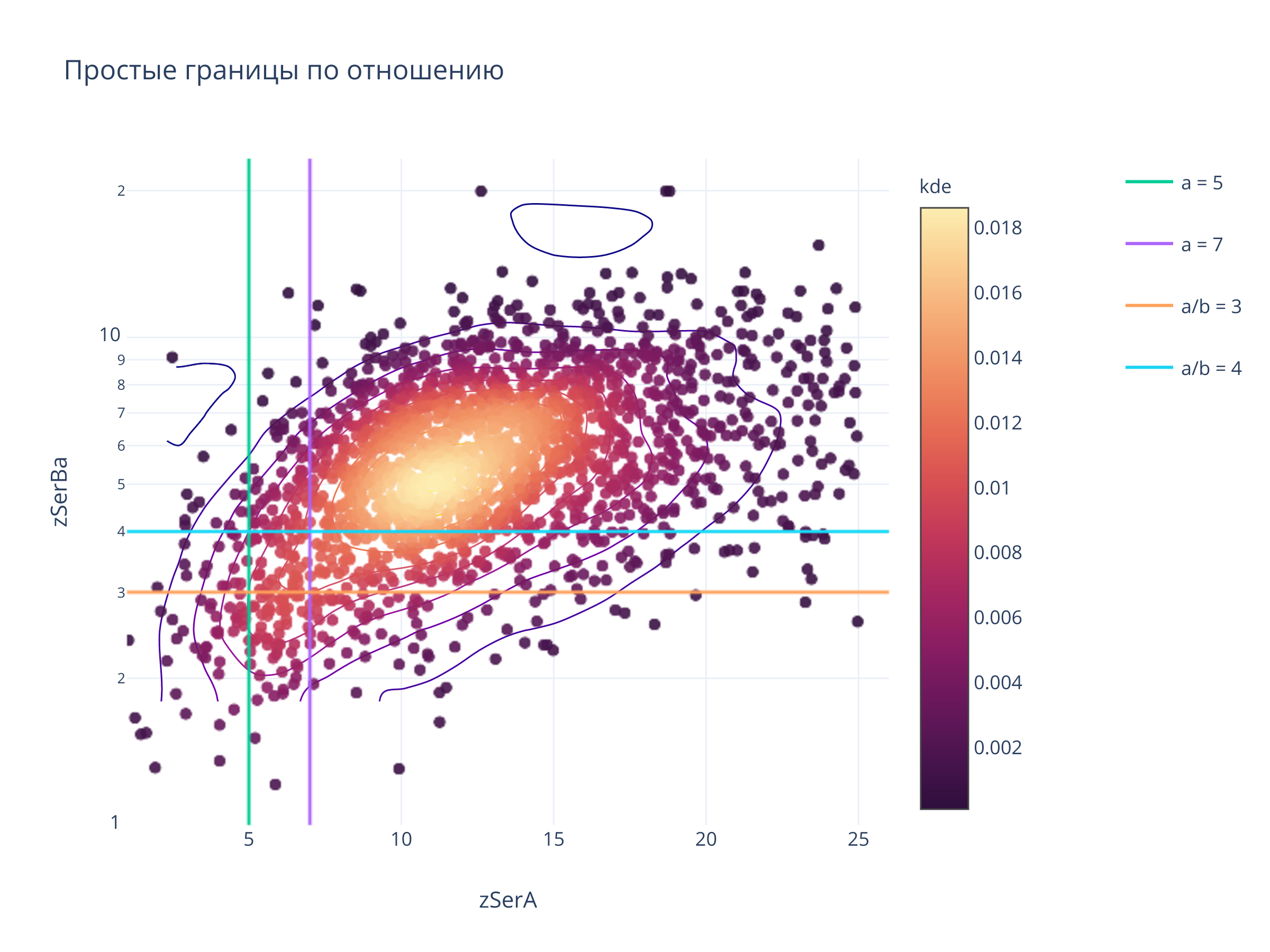
fig.update_layout(
title = "Простые границы по отношению",
clickmode = "event+select",
autosize=False,width=800, height=600)
for x in [5, 7]:
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", name=f"a = {x}", legendgroup=f"a = {x}"))
for y in [3, 4]:
fig.add_trace(go.Scattergl(x = xrange, y = [y,y],mode = "lines",name=f"a/b = {y}", legendgroup = f"a/b = {y}"))
fig
def sel_tresh(t, ratio, amin):
key = f'a/b>{ratio:d} a>{amin:d}'
masks_t[filt][key] = (t[f'{filt}SerA'] > amin) & (t[f'{filt}SerBa'] > ratio)
sqlconds[filt][key] = f"{filt}SerRadius > {amin:.2f} AND {filt}SerAb < {1/ratio:.5f}"
sel_tresh(biggestsermag_aligned, 3, 5)
sel_tresh(biggestsermag_aligned, 3, 7)
sel_tresh(biggestsermag_aligned, 4, 5)
sel_tresh(biggestsermag_aligned, 4, 7)
Покрытие неба¶
fig = go.Figure()
t = biggestsermag_aligned
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
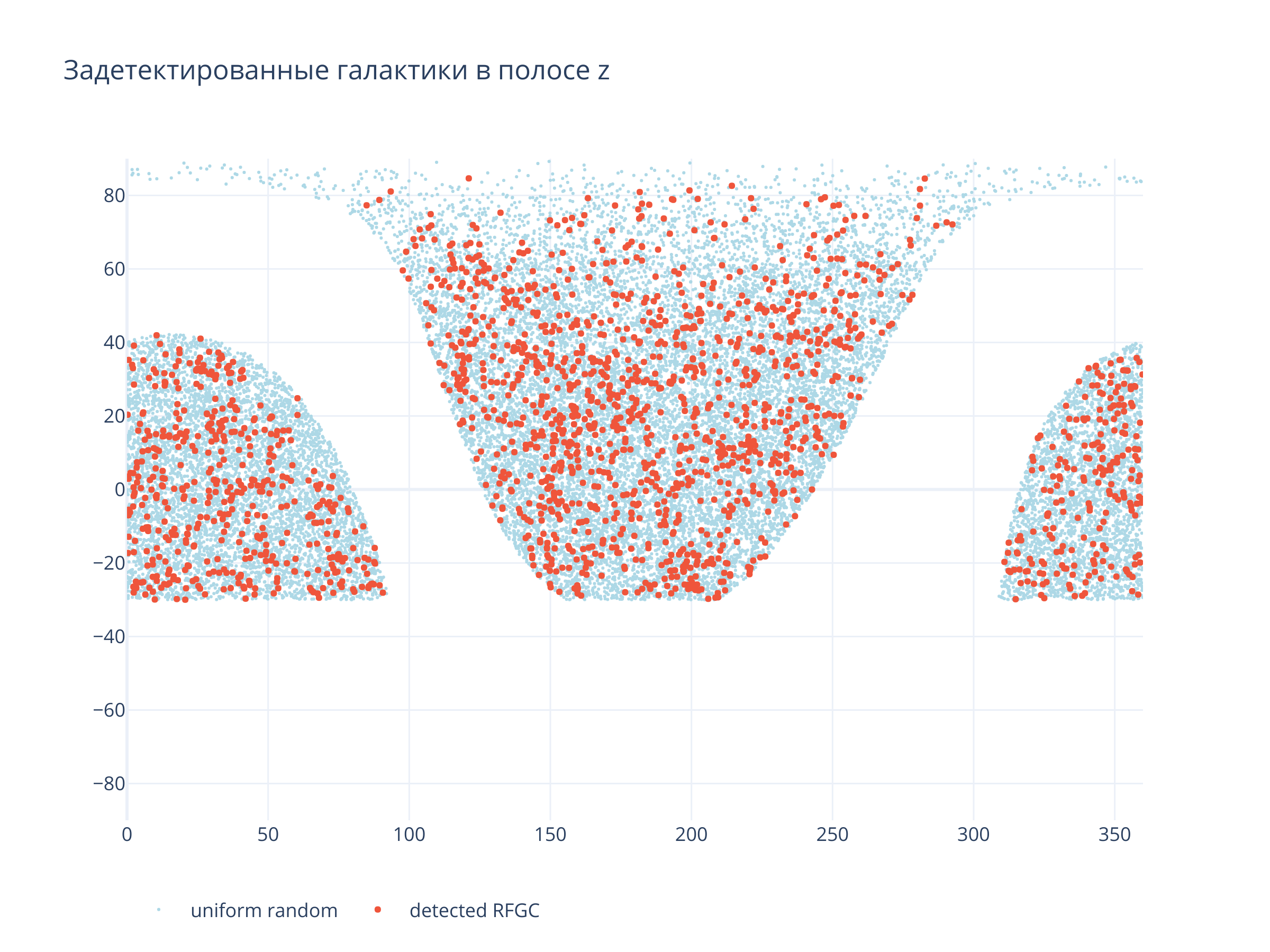
title = f"Задетектированные галактики в полосе {filt}",
legend_orientation = 'h'
)
fig
Светло-синие точки --- равномерно распределённые по области детектирования случайные точки
Выводы¶
filt='z'
summary[filt] = build_summary(biggestsermag_aligned, filt)
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt]
Подбор критериев отбора (y)¶
filt='y'
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex"])
fitted = wshape.dropna(subset=[f"{filt}SerRadius"])
sane = fitted[fitted[f"{filt}SerA"] > 0.051]
aligned = sane[(np.abs(sane[f"{filt}GalPhi"] - sane["PA"]) < 10) & \
(np.abs(sane[f"{filt}SerPhi"] - sane["PA"]) < 10)]
#aligned.index= range(1, len(aligned)+1)
biggestsermag_aligned = aligned.sort_values(["RFGC",f"{filt}SerMag"],ascending=[True, True]).groupby('RFGC').head(1)
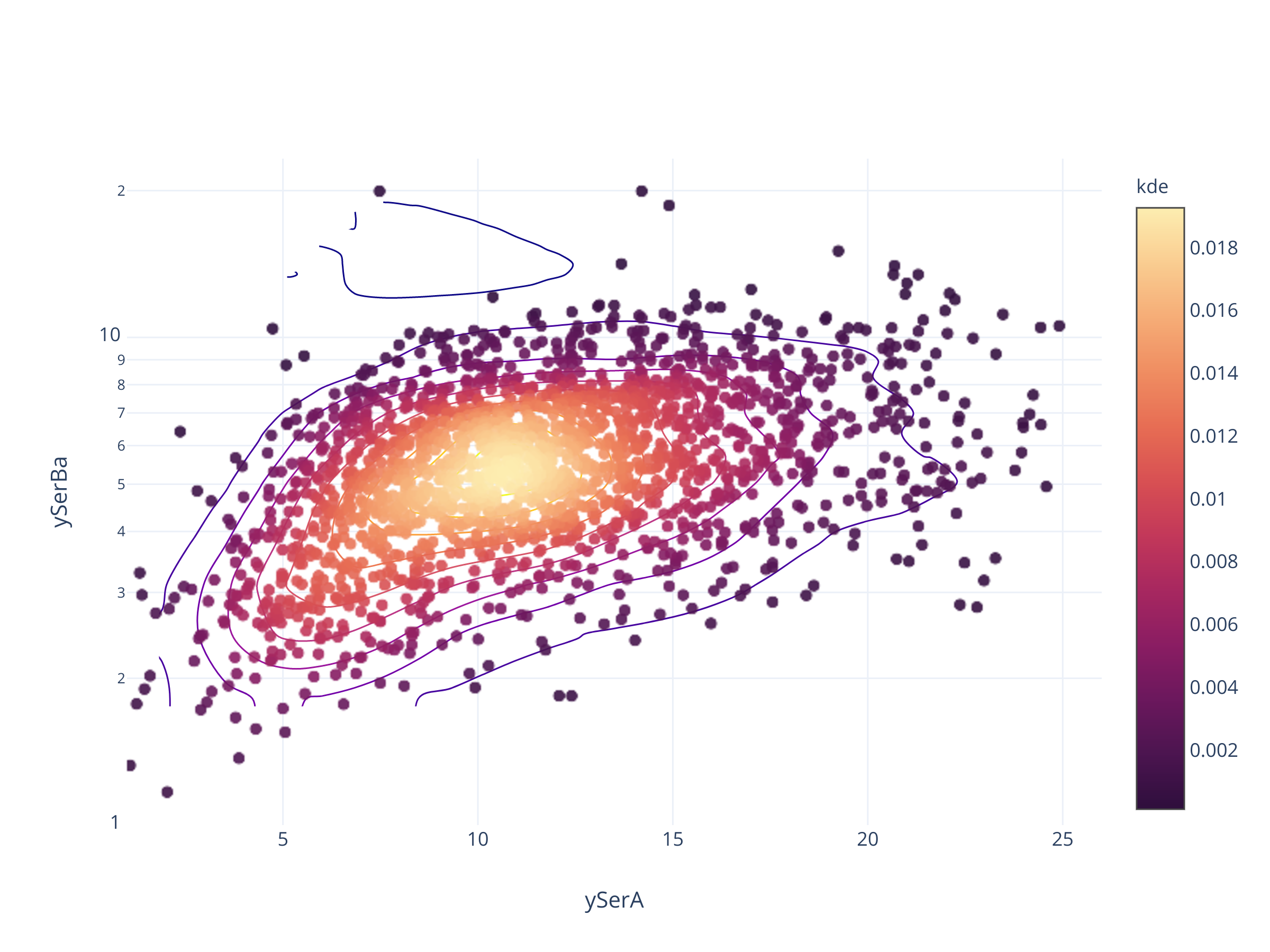
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 1/0.043)
)
fig.update_layout(autosize=False, width=600, height=500)
fig
0. Границы по большой полуоси¶
FWHM у PSF в обзоре $1''-2''$ (Глянул у пары галактик, но надо ещё уточнить)
Если у галактики полуось < $7''$, она будет видна с отношением осей $<7$ (а если нет, то это ошибка алгоритма). Получается, что надо отсекать где-то по $5'' - 7''$
t = biggestsermag_aligned
sizemask = {
5:(t[f'{filt}SerA'] > 5),
7:(t[f'{filt}SerA'] > 7),
}
summary[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index=t.index)}
sqlconds[filt] = {'all':f"{filt}SerRadius > 0.051"}
1. По «наблюдаемому» соотношению осей¶
Можно попробовать подобрать границу «круглости» с помощью этой функции
from code.plotutils import scatter_density_plotly, seaborn_to_plotly
x, y = f'{filt}SerA', f'{filt}SerAb'
y2 = f'{filt}SerBa'
xrange, yrange = (1, 26), (0.043, 1)
t = biggestsermag_aligned
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
rfgc_names = t.loc[mask, 'RFGC']
xx = t.loc[mask, x]
yy = t.loc[mask, y]
sizes = np.linspace(*xrange, 51)
a_b_list = np.array([7, 25])
b_a_list = 1/a_b_list
N = len(a_b_list)
plots_kw = dict(mode="kde", modepars={'bins':(30,30)}, contours=True, alpha=0.9,
pointlabels=rfgc_names)
subplotlayout = dict(rows=1, cols=2)
fig = make_subplots(**subplotlayout)
fig.layout.update(
xaxis = {'domain':[0, 0.4]},
xaxis2 = {'domain':[0.6, 1]},
)
fig.layout.xaxis.type = 'log'
fig = scatter_density_plotly(xx, yy, xrange, yrange, x, y, logy=True, logx=True,
subplotpos={'row':1, 'col':1}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig = scatter_density_plotly(xx, 1/yy, xrange, (1/yrange[1], 1/yrange[0]), x, y2, logy=True, logx=True,
subplotpos={'row':1, 'col':2}, subplotlyt=subplotlayout,
fig=fig, **plots_kw)
fig.update_traces(
marker = {'colorbar':{'x':0.4}},
selector = {'type':'scattergl'},
row = 1, col = 1)
fig.update_traces(
marker = {'colorbar':{'x':1}},
selector = {'type':'scattergl'},
row = 1, col = 2)
fig.update_layout(showlegend=True, legend_orientation="h")
pallete = sb.cubehelix_palette(4, start=.5, gamma=0.8, rot=-0.7, dark=0.25, light=0.75)
ply_chx = seaborn_to_plotly(pallete)
nncurves = len(fig.data)
for i, a_b in enumerate(a_b_list):
fig.add_trace(
go.Scattergl(
x = sizes,
y = b_a_obs(sizes, 1/a_b),
name="%.2f" % a_b, showlegend=True,
legendgroup="%.2f" % a_b,
hoverinfo='name',
line = {'color':ply_chx[i][1]}
), row=1, col=1)
fig.add_trace(
go.Scattergl(
x = sizes,
y = a_b_obs(sizes, a_b),
name = "%.2f" % a_b,
hoverinfo='name', showlegend=False,
line = {'color':ply_chx[i][1]}
), row=1, col=2)
for i, x in enumerate([5, 7]):
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo = "name", legendgroup=f"a = {x}"),
row=1, col=1)
yrange2 = (1/yrange[1], 1/yrange[0])
fig.add_trace(go.Scattergl(x = [x, x], y = yrange2, mode = "lines", line = {'color': ply_chx[i+2][1]},
name=f"a = {x}", hoverinfo="name", showlegend=False),
row=1, col=2)
fig.layout.legend.x = 0.3
fig.layout.legend.y = -0.2
fig.layout.legend.tracegroupgap = 30
fig.layout.title = "Распредение объектов по полуоси и отношению осей и наблюдаемое отношение осей"
fw = go.FigureWidget(fig)
@interact(a_b1 = (1, 30, .1), a_b2 = (1, 30, .1), sigma=(1, 10, 0.1))
def update(a_b1 = 7, a_b2 = 18, sigma=3.5):
with fw.batch_update():
for i, (a_b, inverse) in enumerate([(a_b1, True), (a_b1, False), (a_b2, True), (a_b2, False)],
start=nncurves):
fw.data[i].update(
y = a_b_obs(sizes, a_b, sigma, inverse=inverse),
name = "%.2f" % a_b)
fw.show(height=400, width=900)
def sel_obs_ratio(t, r_obs, sigma, amin):
key = f'r_obs={r_obs:d} sigma={sigma:.2f} a>{amin:d}'
masks_t[filt][key] = (
(t[f'{filt}SerAb'] < a_b_obs(t[f'{filt}SerA'], r_obs, sigma=sigma, inverse=True)) &
sizemask[amin]
)
sqlconds[filt][key] = f"{filt}SerAb < SQRT(SQUARE({filt}SerRadius/{r_obs:.2f}) + {sigma**2:.2f}) / SQRT(SQUARE({filt}SerRadius) + {sigma**2:.2f}) AND {filt}SerRadius > {amin:.2f}"
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 5)
sel_obs_ratio(biggestsermag_aligned, 7, 3.5, 7)
2. По дисперсии линейной регрессии¶
import statsmodels.api as sm
from astropy.modeling import models, fitting
def compute_linregr_pars(t, x, y, xrange, yrange, logx=False, logy=True):
mask=(t[x]).between(*xrange) & (t[y].between(*yrange))
xx_reg = (t.loc[mask, x])
yy_reg = (t.loc[mask, y])
xx_reg = np.log10(xx_reg) if logx else xx_reg
yy_reg = np.log10(yy_reg) if logy else yy_reg
variables = pd.DataFrame({
'const':np.ones_like(xx_reg),
'x':xx_reg,
#'x2':xx**2,
})
linmodel = sm.OLS(yy_reg, variables)
a,b,c,d,e = [0]*5
mod = Namespace()
linresults = linmodel.fit()
mod.eps = linresults.resid
mod.b,mod.a = linresults.params
hist, xe = np.histogram(mod.eps)
xe = (xe[1:]+xe[:-1])/2
ini = models.Gaussian1D()
fitter = fitting.LevMarLSQFitter()
fit = fitter(ini, xe, hist)
mod.sigma_total = fit.stddev
return mod
xrange, yrange = (1, 26), (1, 23.26)
m = compute_linregr_pars(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
x_grid = np.linspace(*xrange, 100)
fitted = 10**(m.a*x_grid + m.b)
upperbnd = 10**(m.a*x_grid + m.b + 2*m.sigma_total)
lowerbnd = 10**(m.a*x_grid + m.b - 2*m.sigma_total)
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
xrange, yrange
)
fig.add_trace(go.Scattergl(x = x_grid, y = fitted, name='fitted'))
fig.add_trace(go.Scattergl(x = np.hstack([x_grid, [x_grid[-1]+1,x_grid[-1]+1], np.flip(x_grid)]),
y = np.hstack([upperbnd, [upperbnd[-1], lowerbnd[-1]], np.flip(lowerbnd)]), line = {'color':'lightgray', 'width':3},
name='2σ range'))
fig.layout.legend.x = 1.3
fig.update_layout(
title = "Линейная регрессия и диапазон в 2σ",
clickmode = "event+select",
autosize=False,width=700, height=500)
fig.show()
def sel_lbnd_linreg(t, amin):
key = f'2 sigma a>{amin:d}'
masks_t[filt][key] = (
(np.log10(t[f'{filt}SerBa']) > m.a*t[f'{filt}SerA']+m.b - 2*m.sigma_total) &
(t[f'{filt}SerA'] > amin)
)
sqlconds[filt][key] = f"-LOG10({filt}SerAb) > {m.a:.5f}*gSerRadius + {m.b:.5f} - {2*m.sigma_total:.5f} AND {filt}SerRadius > {amin:.2f}"
#t = biggestsermag_aligned
sel_lbnd_linreg(biggestsermag_aligned, 5)
sel_lbnd_linreg(biggestsermag_aligned, 7)
3. Простые границы¶
fig = scadensrfgc_ply(
biggestsermag_aligned,
f'{filt}SerA', f'{filt}SerBa',
(1, 26), (1, 23.26)
)
x_grid = np.linspace(*xrange, 100)
fig.layout.legend.x = 1.3
fig.layout.legend.tracegroupgap = 20
fig.update_layout(
title = "Простые границы по отношению",
clickmode = "event+select",
autosize=False,width=800, height=600)
for x in [5, 7]:
fig.add_trace(go.Scattergl(x = [x, x], y = yrange, mode = "lines", name=f"a = {x}", legendgroup=f"a = {x}"))
for y in [3, 4]:
fig.add_trace(go.Scattergl(x = xrange, y = [y,y],mode = "lines",name=f"a/b = {y}", legendgroup = f"a/b = {y}"))
fig
def sel_tresh(t, ratio, amin):
key = f'a/b>{ratio:d} a>{amin:d}'
masks_t[filt][key] = (t[f'{filt}SerA'] > amin) & (t[f'{filt}SerBa'] > ratio)
sqlconds[filt][key] = f"{filt}SerRadius > {amin:.2f} AND {filt}SerAb < {1/ratio:.5f}"
sel_tresh(biggestsermag_aligned, 3, 5)
sel_tresh(biggestsermag_aligned, 3, 7)
sel_tresh(biggestsermag_aligned, 4, 5)
sel_tresh(biggestsermag_aligned, 4, 7)
Покрытие неба¶
fig = go.Figure()
t = biggestsermag_aligned
fig.add_trace(go.Scatter(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scatter(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
title = f"Задетектированные галактики в полосе {filt}",
legend_orientation = 'h'
)
fig
Светло-синие точки --- равномерно распределённые по области детектирования случайные точки
Выводы¶
summary[filt] = build_summary(biggestsermag_aligned, filt)
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt]
Отбор по выполнению критерия в любой полосе¶
bands = list('grizy')
Выберем здесь такие источники, в которые хотя бы в одной полосе вписан Серсик с радиусом выше программного предела
wshape = matches_ref.dropna(subset=[f"{filt}GalIndex" for filt in bands], how='all')
fitted = wshape.dropna(subset=[f"{filt}SerRadius" for filt in bands], how='all')
sane = fitted[(np.any(fitted.loc[:, [f"{filt}SerA" for filt in bands]] > 0.051, axis=1))]
aligned = sane[np.any(
[
(np.abs(sane[galphi] - sane['PA']) < 10) & (np.abs(sane[serphi] - sane['PA']) < 10)
for galphi, serphi in zip(
[f"{filt}GalPhi" for filt in bands],
[f"{filt}SerPhi" for filt in bands])
], axis=0)]
biggestsermag_aligned = aligned.sort_values(
[ "RFGC", *[f"{filt}SerMag" for filt in bands]],
ascending=[True, *[True]*len(bands)]
).groupby('RFGC').head(1)
t = biggestsermag_aligned
ids_per_rfgc = t.groupby('RFGC').count()['objName']
ids_per_rfgc.sort_values(ascending=False)[0:10]
fig = go.Figure()
t = biggestsermag_aligned
fig.add_trace(go.Scattergl(
x = galaxies_sel['ra'],
y = galaxies_sel['dec'], name = "uniform random",
hoverinfo = 'none',
mode = 'markers', marker = {'size': 2, 'color':'lightblue'}
))
fig.add_trace(go.Scattergl(
x = t['RAJ2000'],
y = t['DEJ2000'], name = "detected RFGC",
text = list(t['RFGC']),
hoverinfo = 'text',
mode = 'markers', marker = {'size': 4}
))
fig.update_layout(
xaxis = {'range' :[0, 360]},
yaxis = {'range' :[-90, 90]},
height=800,
title = f"Задетектированные галактики"
)
None
from functools import reduce
def allbandsclause(s, bands="grizy", clause="OR"):
return f" {clause} ".join(["(" + s.format(filt=f) + ")" for f in bands])
def clausejoin(bandkeys, clause="OR"):
return f" {clause} ".join(["(" + sqlconds[k][v] + ")" for k,v in bandkeys.items()])
t = biggestsermag_aligned
sizemask = {
5:np.any([t[f'{filt}SerA'] > 5 for filt in bands], axis=0),
7:np.any([t[f'{filt}SerA'] > 7 for filt in bands], axis=0),
}
filt = 'grizy'
sqlconds[filt] = {'all':clausejoin({k:'all' for k in bands})}
sqlconds[filt]['r_obs a>5'] = clausejoin({
'g':'r_obs=7 sigma=4.00 a>5',
'r':'r_obs=7 sigma=3.80 a>5',
'i':'r_obs=7 sigma=3.50 a>5',
'z':'r_obs=7 sigma=3.50 a>5',
'y':'r_obs=7 sigma=3.50 a>5',
})
sqlconds[filt]['r_obs a>7'] = clausejoin({
'g':'r_obs=7 sigma=4.00 a>7',
'r':'r_obs=7 sigma=3.80 a>7',
'i':'r_obs=7 sigma=3.50 a>7',
'z':'r_obs=7 sigma=3.50 a>7',
'y':'r_obs=7 sigma=3.50 a>7',
})
sqlconds[filt]['2 sigma a>5'] = clausejoin({k:'2 sigma a>5' for k in bands})
sqlconds[filt]['2 sigma a>7'] = clausejoin({k:'2 sigma a>7' for k in bands})
sqlconds[filt]['a/b>3 a>5'] = clausejoin({k:'a/b>3 a>5' for k in bands})
sqlconds[filt]['a/b>3 a>7'] = clausejoin({k:'a/b>3 a>7' for k in bands})
sqlconds[filt]['a/b>4 a>5'] = clausejoin({k:'a/b>4 a>5' for k in bands})
sqlconds[filt]['a/b>4 a>7'] = clausejoin({k:'a/b>4 a>7' for k in bands})
import operator
def maskjoin(bandkeys, action=operator.or_):
return reduce(action, [masks_t[k][v] for k,v in bandkeys.items()])
filt = 'grizy'
masks_t[filt] = {}
masks_t[filt] = {'all': pd.Series(np.ones(len(t), dtype=bool), index=t.index)}
masks_t[filt]['r_obs a>5'] = maskjoin({
'g':'r_obs=7 sigma=4.00 a>5',
'r':'r_obs=7 sigma=3.80 a>5',
'i':'r_obs=7 sigma=3.50 a>5',
'z':'r_obs=7 sigma=3.50 a>5',
'y':'r_obs=7 sigma=3.50 a>5',
})
masks_t[filt]['r_obs a>7'] = maskjoin({
'g':'r_obs=7 sigma=4.00 a>7',
'r':'r_obs=7 sigma=3.80 a>7',
'i':'r_obs=7 sigma=3.50 a>7',
'z':'r_obs=7 sigma=3.50 a>7',
'y':'r_obs=7 sigma=3.50 a>7',
})
masks_t[filt]['2 sigma a>5'] = maskjoin({k:'2 sigma a>5' for k in bands})
masks_t[filt]['2 sigma a>7'] = maskjoin({k:'2 sigma a>7' for k in bands})
masks_t[filt]['a/b>3 a>5'] = maskjoin({k:'a/b>3 a>5' for k in bands})
masks_t[filt]['a/b>3 a>7'] = maskjoin({k:'a/b>3 a>7' for k in bands})
masks_t[filt]['a/b>4 a>5'] = maskjoin({k:'a/b>4 a>5' for k in bands})
masks_t[filt]['a/b>4 a>7'] = maskjoin({k:'a/b>4 a>7' for k in bands})
summary[filt] = {}
Выводы¶
summary[filt] = build_summary(biggestsermag_aligned, 'grizy')
Таблица¶
Здесь:
| N | сколько отобралось объектов, соответствующих галактикам RFGC по критерию |
| N/N0 | доля всего каталога RFGC |
| N/Nd | c поправкой на неполное покрытие неба |
| PS | нашлось в PanSTARRS по критерию |
summary[filt].loc['r_obs a>5', 'PS'] = 13849922
summary[filt].loc['r_obs a>7', 'PS'] = 7492787
summary[filt].loc['2 sigma a>5', 'PS'] = 15569760
summary[filt].loc['2 sigma a>7', 'PS'] = 8922280
summary[filt]
Почему тут для всего вместе получается больше 1 я не знаю. Возможно, так случайно получилось из-за неравномерности распределения галактик.